The future of data storage lies in the foundation of biology
DNA holds the key to storing vast amounts of digital data
 DNA (source: Pixabay)
DNA (source: Pixabay)
When we think of DNA, we inevitably think of genes and the biological instructions that our bodies follow to produce—well—us. DNA is a language composed of As, Ts, Gs, and Cs whose pattern dictates the characteristics that each and every one of us uniquely possess. But the interesting thing about DNA is that its data storage capability isn’t necessarily limited to biological uses.
In fact, what do deoxyribonucleic acid and magnetic tape have in common? They can store digital data in a similar way, albeit with different materials and therefore different outcomes.
Magnetic Tape as a Means of Storing Digital Data
Magnetic tape was first introduced in 1952 with IBM’s 762 tape. Since then, the size of magnetic tapes has drastically been reduced while still increasing storage capacities significantly. The 2012 Linear Tape-Open (LTO) 6, for example, took data storage up to 2.5 TB. Compare this with the original IBM 762 tape, which stored only 2 million digits and was 1,200 feet in length!
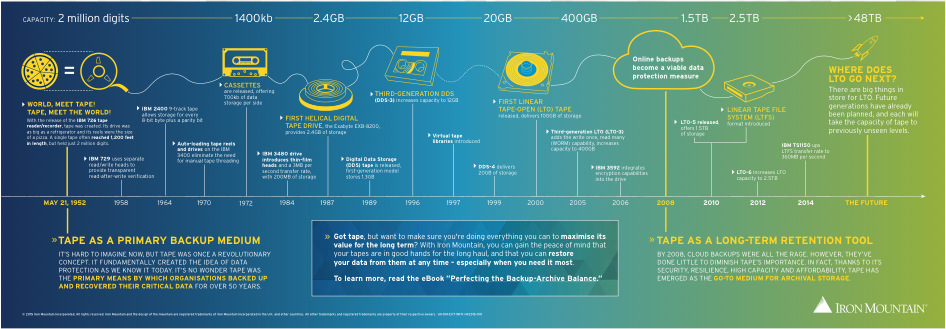
The process of recording information on magnetic tape utilizes the binary system of 0s and 1s, which correspond to two magnetic polarities of the tape. A current-carrying conductor, known as the write head, can detect the magnetic quality of the tape and alter it according to the needed pattern. In contrast, the read head can produce current from the magnetic pattern, recording the binary code and thereby decoding the stored data.
Recording data on magnetic tape continues to be a viable option but two major issues exist with this form of storage:
-
The world’s data is doubling every two years, making it difficult for magnetic tape production to keep up with such a large quantity of information
-
According to studies, current magnetic tape has an attrition rate of 22% and only lasts 5 to 10 years before they must be replaced to preserve the stored data
The Difference in DNA Data Storage
There are currently 50 trillion trillion trillion base pairs of DNA on planet Earth; that’s 50 billion tons of DNA in total and good evidence as to the amount of information stored in this biological technology, considering that it comprises every living being.
DNA is a very dense form of storage. One gram of the substance could potentially hold a billion gigabytes of information, in a form factor that will well outlast any current data storage material. With evidence from the known fossil record, DNA provides a promising unit to hold information for hundreds of thousands of years, given the appropriate preservation conditions.
For example, genetic material found in fossils of plants and insects recovered from Greenland ice cores dates to 800,000 years ago. The absence of oxygen and water mean that DNA can be better preserved and persist from ancient times to the contemporary world.
Natural methods of preservation imply positive results for digital storage in genetic material, but scientific advances in the glass encasing of DNA provide even greater hope for long-term use. The double-stranded integrity of DNA is preserved after being encapsulated by sol-gel derived silica, according to a paper by Kapusuz and Durucan. With novel methods to protect the biological matter, there is every reason to consider it as an alternative digital storage material.
However, the general approach of encoding information in DNA brings up a relevant issue: what is the cost of this type of data storage and how does it compare to the costs of current storage methods? According to synthetic DNA manufacturer, Twist Bioscience, the cost of storing information in DNA is equivalent to the costs of the reagents needed to add additional bases to DNA molecules. Twist’s estimation of these costs is “less than a fraction of a penny per gigabyte to encode and recover stored information” (DNA-Based Digital Storage). Assuming the synthetic DNA would possess a longer shelf-life, the cost of this data storage method is cheaper than the alternative magnetic tape option.
Of course, the cost of sequencing or “reading” the DNA must also be factored in. The human genome project from 1990–2003 required roughly $3 billion to sequence the entire human genome, about 4 GB of information. In 2014, the same process could be done for about $1,000 due to Illumina’s whole-genome sequencing efforts. That represents a huge cost reduction, which provides hope for greater affordability in the future, despite the fact that storing 1 TB of information on a disc drive currently costs $100.
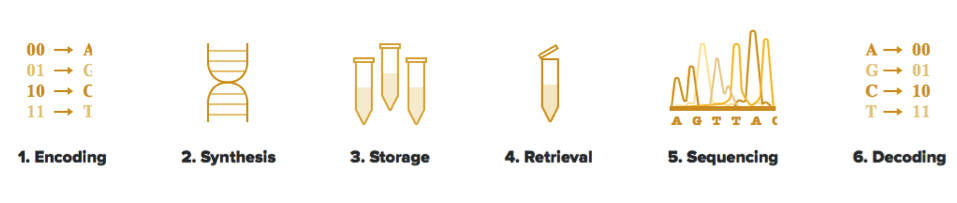
The process of storing information in DNA is straightforward in that the binary code traditionally used in information technology is converted to the equivalent adenine, thymine, guanine, and cytosine nucleotides that make up genetic material. This conversion is acquired through algorithms from telecommunications and other information technologies, according to Twist Bioscience. The company provides the following example of a conversion between binary code and nucleotide bases: “00 represents A, 10 represents C, 01 represents G, and 11 represents T.” Increasing location choices from two (0 or 1) to four (A, T, G, or C) is an indication of an increase in efficiency from Twist’s perspective.
But what about errors in media transmissions? Algorithms known as error correction codes have been developed to prevent error transmissions in the transition between base two and base four codes, that is between binary code and DNA bases. The error correction codes are the primary reason for the short length of approximately 100 to 200 bytes of the then synthesized DNA strands. These chunks of DNA must be ordered correctly with the help of “biological barcodes” that are created at the ends of the synthesized segments (Twist Bioscience).
To determine the success of the encoding process, these strands of DNA can undergo reverse processes, which return the genetic material to binary code that can be read as the original digital information. This involves isolating the DNA and amplifying it through a common laboratory technique, the polymerase chain reaction (PCR). The DNA can then be sequenced in short chunks and once the base pattern is established, it can be reconverted to binary code and pieced together to obtain the initial digital information. Because a portion of the DNA is taken for sequencing, each strand must be duplicated through PCR to ensure that the original data is still available, even after it is decoded.
The time is takes to decode information in DNA is another measure of its feasibility for use as a storage medium. In 2014, the human genome could be sequenced at a rate of approximately 1 GB per hour. At the same time, Illumina could sequence 18,000 human genomes per year, translating to about 72 TB of information (to put 72 TB into perspective, that’s about 24 disc drives, each roughly the size of a book, with 3TB of storage per drive).
With the advent of new technology, again by Illumina, 4 GB of DNA sequencing can now be accomplished in one hour!
Current Advances in DNA Data Storage
DNA storage is not a new concept. In 2012, scientist George Church and colleagues at Harvard University encoded Church’s book Regenesis: How Synthetic Biology Will Reinvent Nature and Ourselves.
Since then, other advancements have been made in technology which have established genetic material as an even more relevant material for digital information storage. In a recent partnership between Twist Bioscience, Microsoft, and the University of Washington, performances of two musical pieces at the Montreux Jazz Festival have been successfully stored in DNA using the process described above.
DNA storage is meant to condense and replace magnetic tape storage, miniaturizing everything in the process and therefore saving space for the large quantities of data we produce. Microsoft is thought to have the goal of including DNA in the cloud as well as having genetic material in data centers for operational storage.
Besides the storage longevity and compactness of DNA, there remains an important point about the medium. Deoxyribonucleic acid is a substance from nature itself. Technology is headed in the direction of biomimicry and therefore what storage element would be more fitting to house the vast quantities of digital information we create than the storage capacity that supports virtually all life on Earth?
