 DNA Lab. (source: University of Michigan School of Natural Resources & Environment on Wikimedia Commons)
DNA Lab. (source: University of Michigan School of Natural Resources & Environment on Wikimedia Commons) Three years ago, we set out to make personal DNA tools that everyone could use. miniPCR started as a maker project, and after a design and prototyping phase, we launched the DNA Discovery System on Kickstarter in 2014. We exceeded our funding goal threefold, which enabled us to scale up production and mount distribution logistics. We have since built a team of biologists, designers, and engineers that operates out of the Harvard Innovation Launch Lab, a community of entrepreneurs and innovators on Harvard University’s dynamic Allston Campus. Manufacturing operations are located just outside of Boston, from where we’ve now shipped DNA Discovery Systems to hundreds of researchers, educators, and DNA curious individuals in every continent.
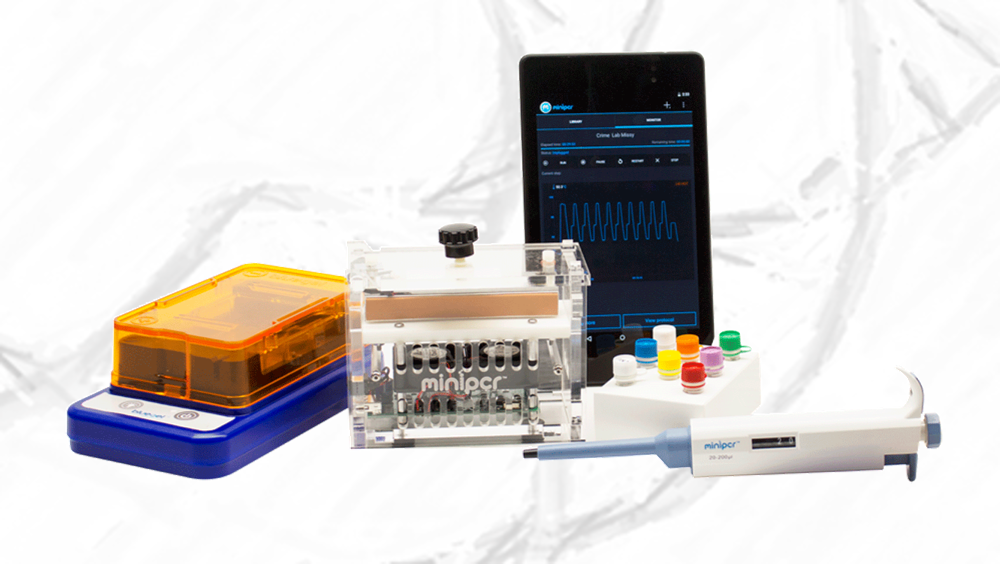
The miniPCR DNA Discovery System is a portable, personal DNA lab that anyone can use. It contains the fundamental tools of DNA analysis: a PCR thermal cycler, a gel electrophoresis apparatus with a built-in blue-light transilluminator, and a variable volume micropipette. It is portable, user friendly, operable through smartphones or computers, and is priced at $990. Its components are also available individually so users can build-to-fit their DNA labs. For example, lab-based researchers may need greater PCR capacity and educators may chose to start with the simpler techniques of DNA gel electrophoresis and visualization. Field-based users may opt to power miniPCR with rechargeable batteries, and couple it to downstream sequencers foregoing gel electrophoresis components. The miniPCR DNA Discovery System (pictured in Figure 1-1) is truly a modular ecosystem that can be configured to the user’s needs.
When we first shared our story with the Biocoder community (fall 2014 issue), we didn’t know the speed at which miniPCR and the field in general would move. We started miniPCR driven less by a specific goal than by sheer curiosity: what would the world look like if everyone had access to DNA science? What would people use miniPCR for? We were excited about enabling new possibilities, about what might happen if we removed barriers to experimentation, expanding channels for scientific innovation. Over the last three years, miniPCR has been in the hands of hundreds of new users. In this article, we share the stories of some of our inspiring users to illustrate the explosion of creativity and innovation that they are leading. There are many more stories that we’d like to share, but just won’t fit the page.
Reimagining Animal Health
Imagine you’re a pig farmer. You live on a small farm, in the Philippines. Your animals are your family’s sole source of income, as long as they’re healthy.
You know that any day one of your pigs can catch the flu–the swine flu. Living in tight quarters, one pig coughing and sneezing can quickly lead to the next pig coughing and sneezing, until an outbreak of swine flu has taken over your farm. If it’s a bad enough virus, the health of your herd can be gone in the blink of an eye. If you called in a veterinarian, she would visit your farm, take samples from your pigs’ noses and mouths, and drive back to the city to analyze those samples in the lab. Two weeks later, you’d learn the results. Two weeks might be just the time it takes for infection to spread and take away your way of life.
But it doesn’t have to be that way. Today farmers can collect those samples themselves. They can jump right into the pen and swab their pigs’ mouths with a little filter paper. Then, they can take those paper swabs, place them in a test tube, and mix them with chemicals that help isolate genetic material from their pigs’ mouths. Without leaving their farms, they can place a drop of genetic material into a little analyzer smaller than a shoebox, program it to detect DNA or RNA from the flu virus, and within one hour farmers can visualize the results.
A Personal DNA Revolution
This reality is possible because we’re living in an era of personal DNA technology (Figure 1-3). Several groups around the world are working to put DNA technology in the hands of everyday people. This movement has been defined under different labels: biohackers, do-it-yourself (DIY) biologists, citizen scientists, etc. Behind these labels lies a common direction of change: through accessible DNA analysis tools, every one of us can test and tinker with DNA ourselves, in our basements and kitchens.

DNA is strung into a long and twisted double helix that in humans has a total of three billion base pairs beginning to end. The lines that carry genetic information, also known as genes, can be as short as a few dozen to several thousand base pairs long. So to find useful genetic information, such as DNA that can answer, “Is my pig sick?” we typically don’t need to read all three billion bases. That would be like craving pizza at night and having to read the phone book from cover to cover, pausing at every line, in search for the nearest pizza joint.
Three decades ago, humans started to invent tools that can very quickly find a specific chunk of genetic information, no matter how small. In a test tube, they can find any given line of DNA. But that DNA is still very small and surrounded by a lot of other DNA. In order to make that specific gene stand out above the rest, we can make copies of the gene, in a process known as the polymerase chain reaction (PCR). PCR machines (also known as thermal cyclers or thermocyclers) can very quickly make billions of copies of a target DNA sequence, and the copies accumulate until we can finally see that target piece of DNA, detect it, read it, and understand it (Figure 1-4). The process is a way to answer questions not only about swine flu in pigs but also those encoded in our own DNA: Am I at risk for ovarian cancer? Am I of Irish descent? Is this child my son?
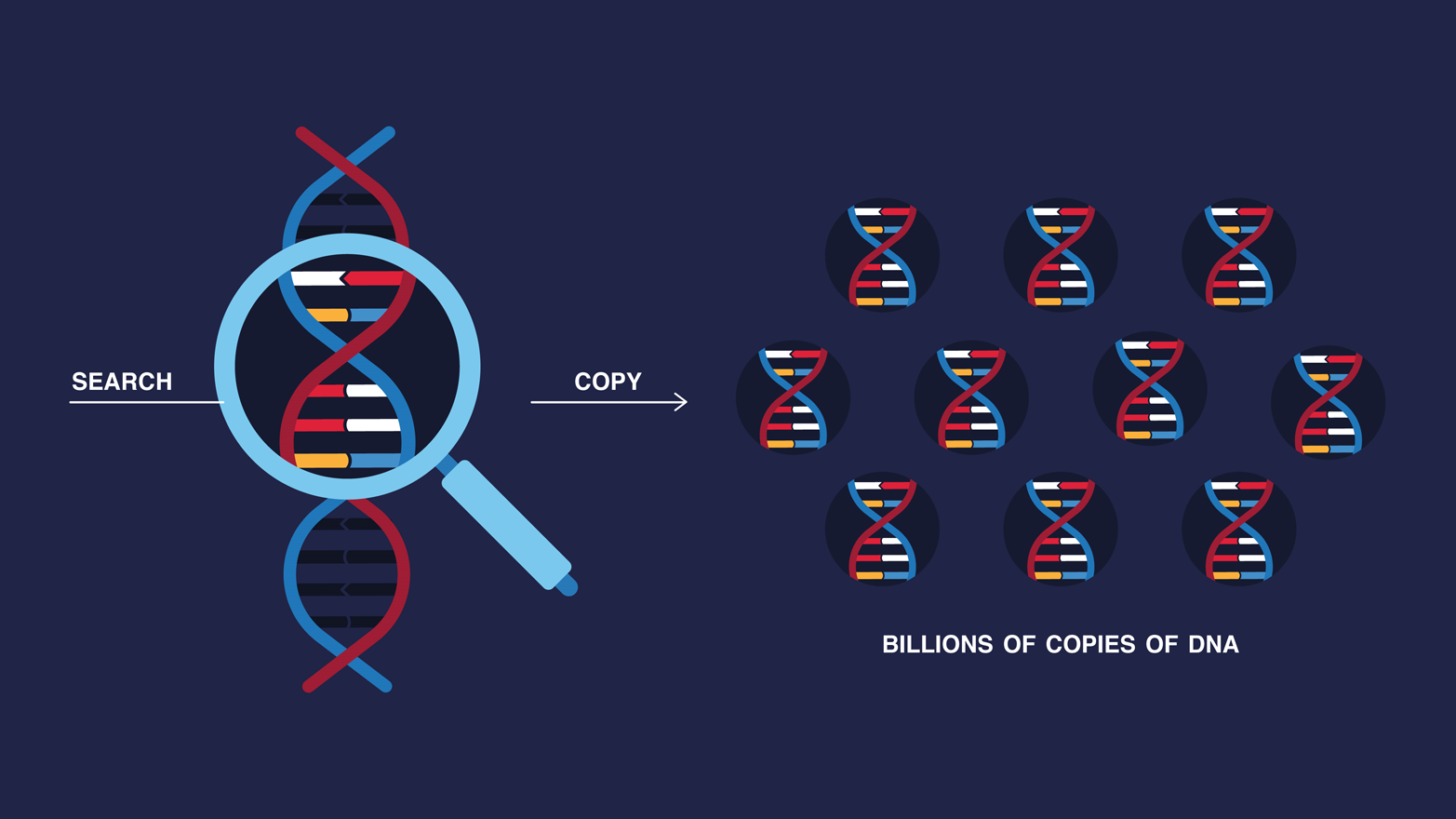
This ability to quickly make millions of copies of DNA by PCR, as simple as it sounds, has transformed our world. PCR machines help scientists detect disease, modify crops, decide when food is safe to eat or when it’s contaminated with deadly bacteria. They help judges decide who goes to jail and who is released based on DNA evidence. For this invention, Kary Mullis was awarded the Nobel Prize in Chemistry in 1993.
Today, this power is accessible to every one of us, as PCR and other DNA technologies become increasingly available outside of professional labs.
The Truffle Farmer That Needed DNA Analysis
People often ask us where this revolution in accessibility might take us, or how the world will change. However, it is notoriously hard to predict how new technology will transform society. For example, when we first developed miniPCR, we could not have predicted that a truffle farmer would have a use for DNA analysis. Dr. Paul Thomas grows truffles for a living, and we see him pictured in Figure 1-5 holding the first UK-cultivated truffle.
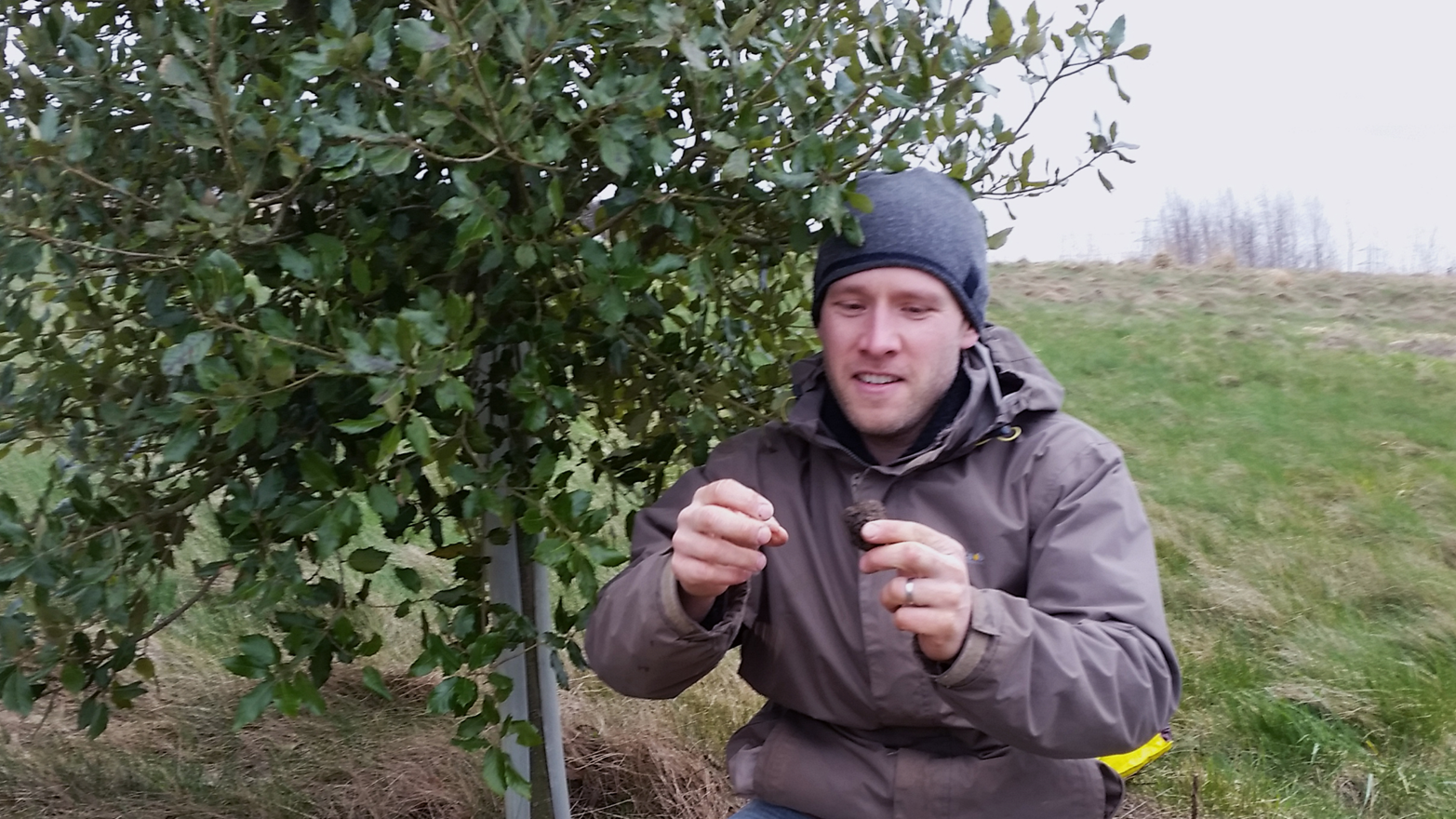

Truffles are a delicacy that stems from a fungus growing on the roots of living trees, and it’s a really rare fungus: some species may fetch between $3,000 to $7,000 per kilogram, or even more. We’ve learned from Thomas that the stakes for a truffle farmer can be really high: When he sources new truffles to grow on his plantations, Thomas is exposed to knock-offs that even to an expert’s eye can look and feel like the real thing. Even under a microscope, these lower quality truffle spores can look authentic. So in order to select the right truffles to grow on his farm, the ones that chefs all around the world will fight over, Thomas has to use DNA analysis. This is mind-blowing, and it’s hard now for us to look at a black truffle risotto without thinking of its genes.
Conservation Biology: Taking the DNA Lab into the Field
Personal DNA technology also means that professional scientists can now take their labs into the field. Elaine Guevara is pursuing her PhD in anthropology from Yale. She’s interested in primate evolution and spends significant time in Southwestern Madagascar, studying a colony of sifaka lemurs on the Beza Mahafaly Reservation (Figure 1-6).
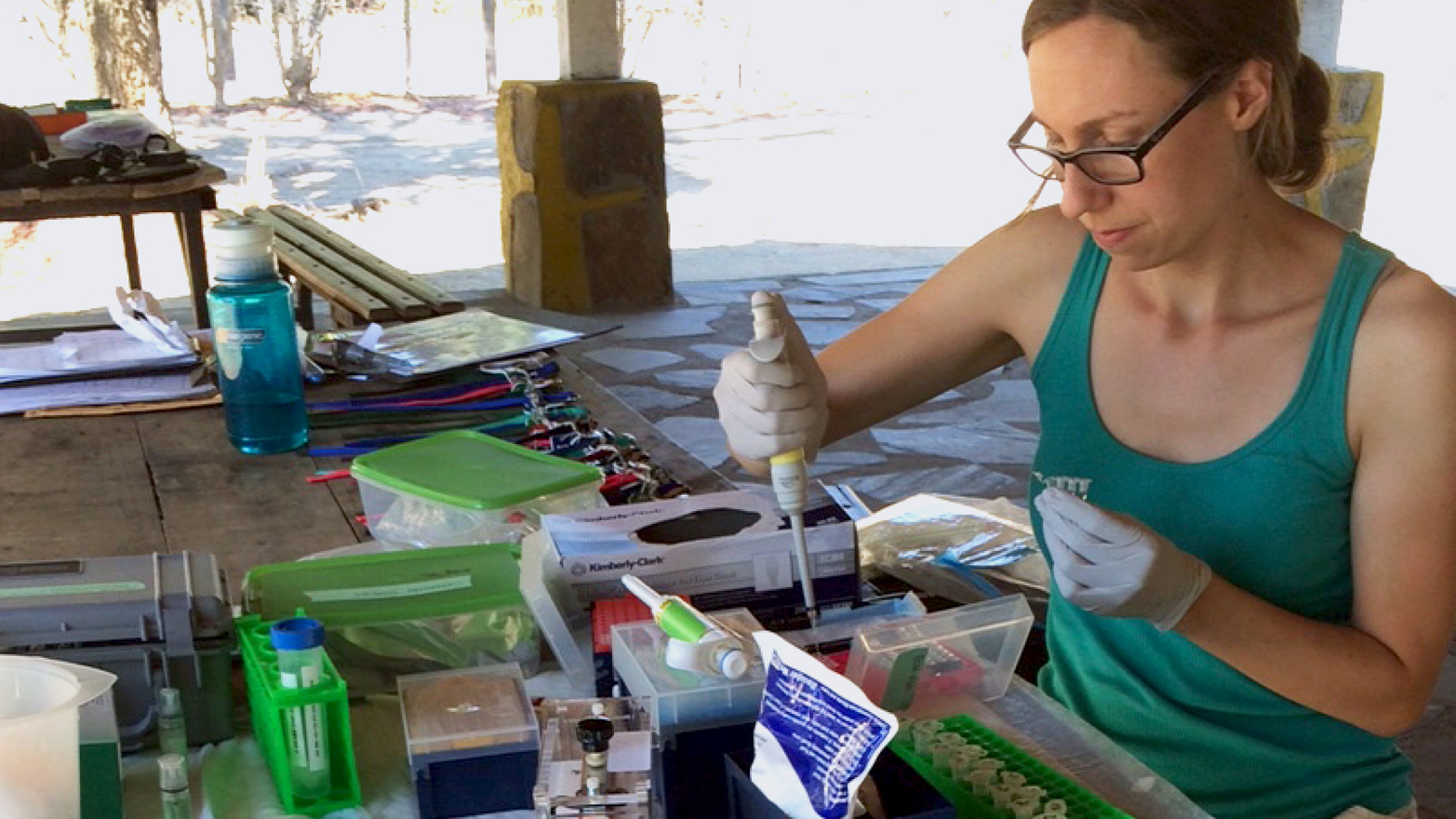

This colony has only 700 lemurs left. So few that these lemurs now share a lot of genes in common. And if they keep inbreeding, they may face health risks or even extinction. To help prevent that, Guevara is monitoring their genetic diversity; but getting a bunch of endangered lemurs on a plane so you can study their genes back at Yale is not really a viable option—imagine the hassle at customs. With miniPCR, Guevara can now make billions of copies of these lemurs’ genomes right in the Madagascar forest, powered by a simple solar battery, and bring back to her lab in the US just test tubes that contain the lemurs’ complete genetic information via whole-genome amplification (WGA).
Saving Lives: Ebola Detection in Makeni, Sierra Leone
We can also use personal DNA technology to save human lives. In 2015, Cambridge University professor Ian Goodfellow traveled to Sierra Leone to try and diagnose patients with Ebola . Doctors there were limited in what they could do; test results could take more than a week to come back, by which time it was too late for those suffering from the disease—and their relatives.
So Goodfellow decided to move his lab to Makeni—that’s him with 10 tons of equipment in the picture—and set it up in a pop-up tent which when fully functional, would be able to turn samples around in 24 hours or less. Only one problem: the high-end DNA analysis equipment didn’t work like it would under controlled lab conditions in the UK. We’re talking 35°C heat and over 90% humidity in Makeni. So instead, he used miniPCR machines, small enough to be placed in front of the air conditioning unit, to keep the virus sequencing workflow going, and keep saving lives.
Genes in Space: DNA Analysis Beyond Earth
Goodfellow’s pop-up lab in Makeni may seem like an extreme place for DNA testing, but let’s travel to an even more extreme place, beyond Earth. Let’s talk about DNA analysis in outer space. Astronauts live aboard the International Space Station, 250 miles above the Earth, orbiting the planet at 17,000 miles per hour, so they often see 15 sunsets and sunrises every day. They live in microgravity, where human bodies can start to do weird things. For example, when astronauts are in space for an extended period of time, their immune systems get weakened, making them more prone to infection.
Why do astronauts experience immune system suppression? Could epigenetic changes be the cause? These were the questions that a high-school student from New York, Anna-Sophia Boguraev, wanted to answer. Anna-Sophia is one of hundreds of students who have participated in Genes in Space, a science competition organized by miniPCR and partners (Boeing, the Center for the Advancement of Science in Space, New England Biolabs, and Math for America).
Anna-Sophia designed a DNA experiment that uses miniPCR to test her hypotheses about immune suppression aboard the International Space Station. On April 8, 2016, miniPCR and Anna-Sophia’s experiment were launched to space from NASA’s Kennedy Space Center in Cape Canaveral. Figure 1-8 shows Anna Sophia watching her experiment launch, and the stream of smoke is indeed the rocket that brought miniPCR to the International Space Station.
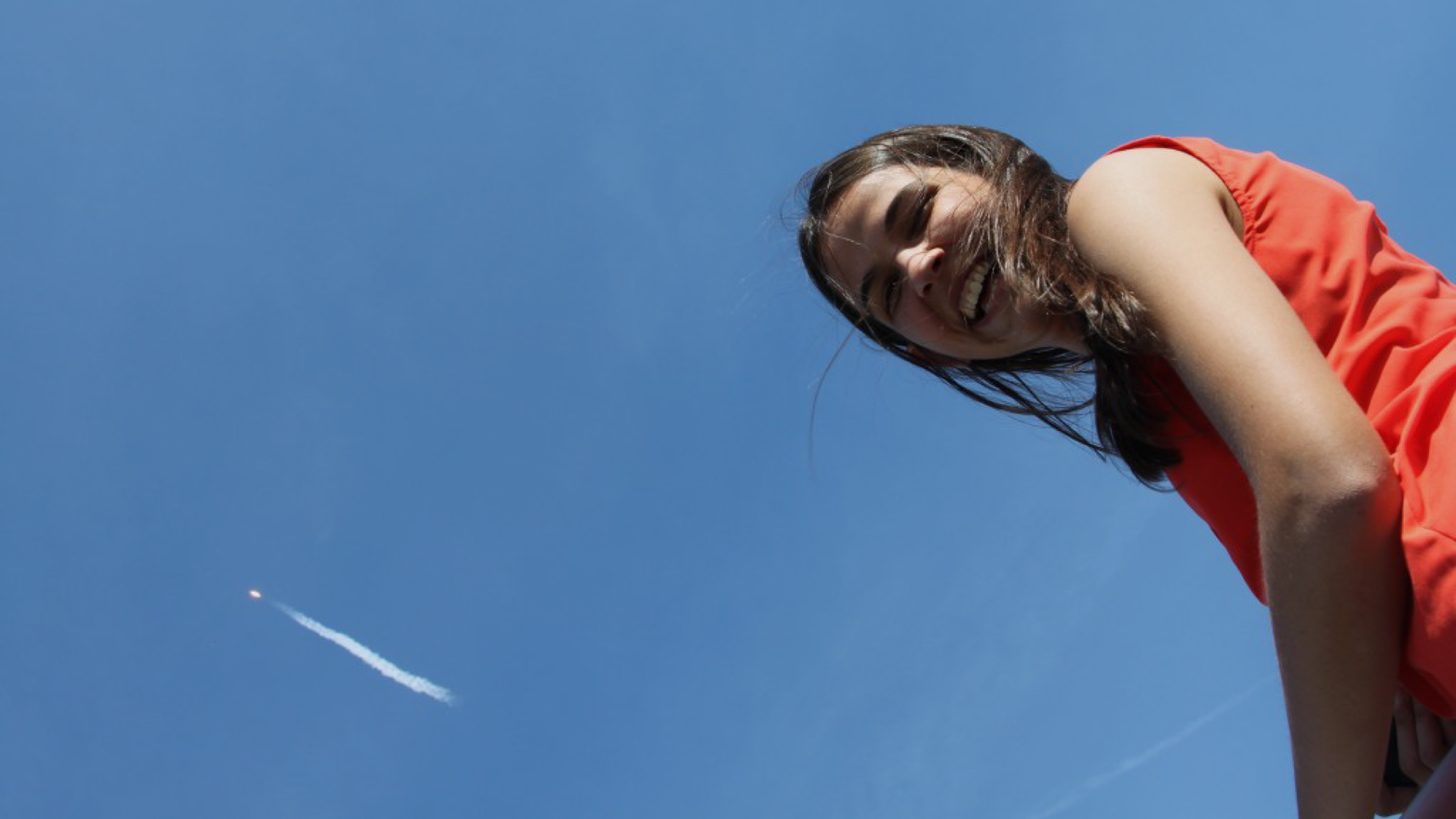

A few days later, astronauts carried out the first DNA amplification experiment in space, and British astronaut Tim Peake conducted Anna-Sophia’s experiment aboard the Space Station (Figure 1-9). We are now in the process of analyzing the results (we know that PCR in space was successful!). miniPCR is now aboard the International Space Station, where it can help monitor living conditions and protect the lives of astronauts. Its efficient design allowed miniPCR to open up a new era of genetics in space.

DNA Knocking on Everyone’s Doors
A 17-year-old student designing a DNA analysis experiment to protect the lives of astronauts may sound like a rarity, or the mark of a child genius. But to us, this signals something bigger, that DNA science is finally becoming within the reach of everybody. Just like a few years ago, a college student armed with a personal computer could code an app in his dorm, an app that is now a social network with more than one billion users.
Could we be moving into a world of one personal DNA machine in every home? We already work with families that are living in such a reality. The Daniels family, for example, set up a DNA lab in the basement of their suburban Chicago home last summer (Figure 1-10). This family isn’t made up of PhD scientists. Instead, they are like many other families who are looking for fun, creative ways to spend time together.

In fact, the dad of this family, Bryan, said all he wanted to do was to “keep his kids off the XBox this summer,” so he replaced the Xbox for the personal DNA box. By day, Bryan is an executive at a private equity firm. On nights and weekends, he plays with miniPCR alongside his kids aged 7 and 9, as a way to explore the living world. Last time I spoke with them, they were testing home-grown produce from their backyard garden. They had picked fresh tomatoes, mixed them with a simple DNA extraction solution, and with their home DNA copier, they were ready to find out whether these tomatoes had genetically engineered traits. For them, the DNA Discovery System is the equivalent of the chemistry set for the 21st century.
Are You DNA Curious?
In 2016, most of us may not yet be diagnosing genetic conditions in our kitchen sinks, or doing at-home paternity testing, but we’ve reached a point in history where you, working in your own kitchen, can take DNA into your own hands and copy it, paste it, analyze it, and extract meaningful information from it.
Explosive innovation is bound to happen. A powerful, transformative technology that was only available to a select few in the ivory tower can now be in your hands, and in the hands of everyone, from farmers to school children. Think about when telephones stopped being attached to the wall via cords, or when computers left the mainframe and entered everyone’s offices and homes.
The ripples of this personal DNA revolution may be unexpected. But here’s the thing about revolutions: they don’t go backward. Access to DNA technology is spreading faster than our imagination. So if you’re curious, get up close and personal with DNA today. It’s in our DNA to be curious.