Deep learning meets genome biology
An interview with Brendan Frey about realizing new possibilities in genomic medicine.
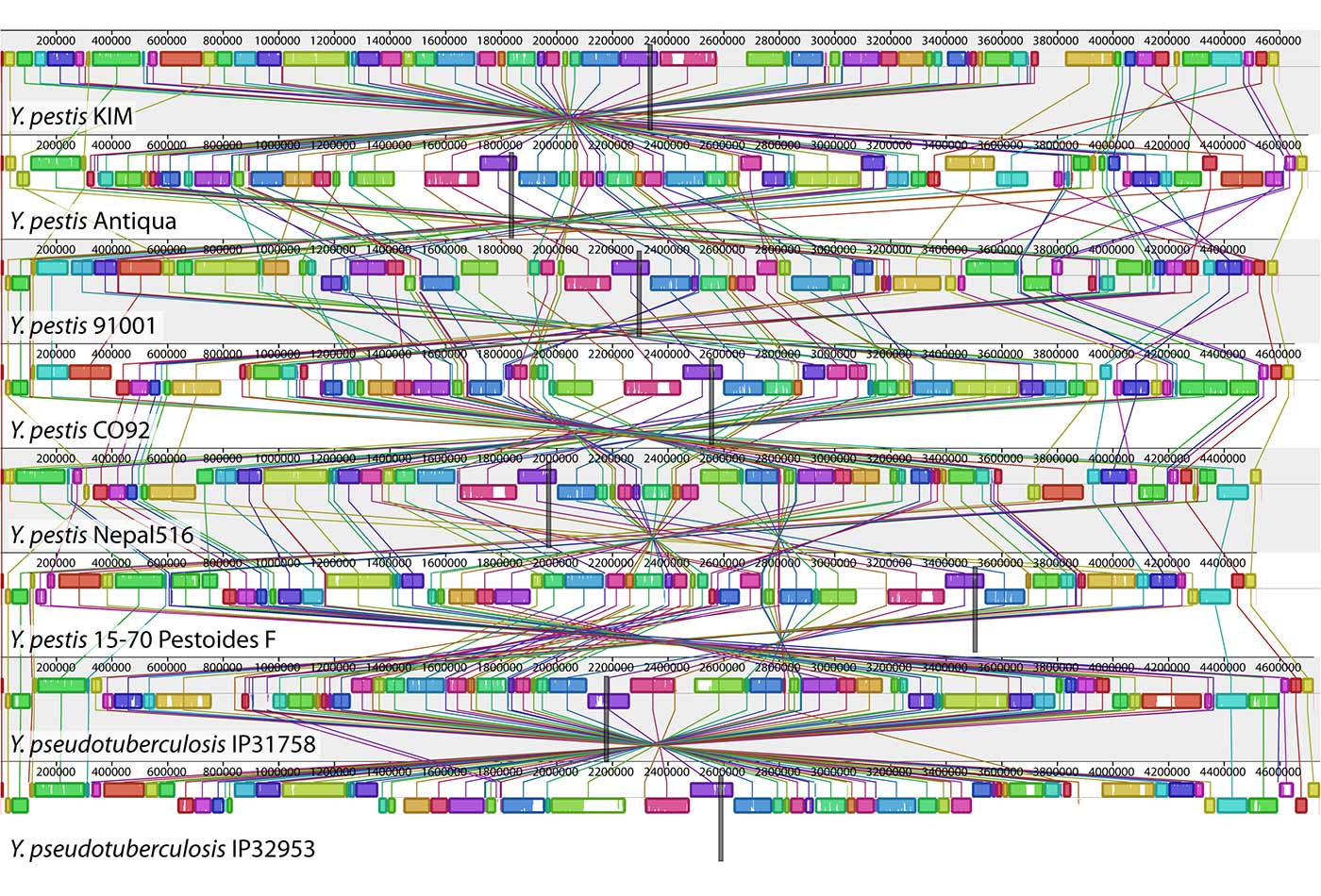 A genome alignment of eight Yersinia isolates. (source: PLOS Genetics on Wikimedia Commons)
A genome alignment of eight Yersinia isolates. (source: PLOS Genetics on Wikimedia Commons)
As part of our ongoing series of interviews surveying the frontiers of machine intelligence, I recently interviewed Brendan Frey. Frey is a co-founder of Deep Genomics, a professor at the University of Toronto and a co-founder of its Machine Learning Group, a senior fellow of the Neural Computation program at the Canadian Institute for Advanced Research, and a fellow of the Royal Society of Canada. His work focuses on using machine learning to understand the genome and to realize new possibilities in genomic medicine.
Key Takeaways
- The application of deep learning to genomic medicine is off to a promising start; it could impact diagnostics, intensive care, pharmaceuticals and insurance.
- The “genotype-phenotype divide”—our inability to connect genetics to disease phenotypes—is preventing genomics from advancing medicine to its potential.
- Deep learning can bridge the genotype-phenotype divide, by incorporating an exponentially growing amount of data, and accounting for the multiple layers of complex biological processes that relate the genotype to the phenotype.
- Deep learning has been successful in applications where humans are naturally adept, such as image, text, and speech understanding. The human mind, however, isn’t intrinsically designed to understand the genome. This gap necessitates the application of “super-human intelligence” to the problem.
- Efforts in this space must account for underlying biological mechanisms; overly simplistic, “black box” approaches will drive only limited value.
David Beyer: Let’s start with your background.
Brendan Frey: I completed my Ph.D. with Geoff Hinton in 1997. We co-authored one of the first papers on deep learning, published in “Science” in 1995. This paper was a precursor to much of the recent work on unsupervised learning and autoencoders. Back then, I focused on computational vision, speech recognition, and text analysis. I also worked on message passing algorithms in deep architectures. In 1997, David MacKay and I wrote one of the first papers on “loopy belief propagation” or the “sum-product algorithm,” which appeared in the top machine learning conference, the Neural Information Processing Systems Conference, or NIPS.
In 1999, I became a professor of computer science at the University of Waterloo. Then in 2001, I joined the University of Toronto and, along with several other professors, co-founded the Machine Learning Group. My team studied learning and inference in deep architectures, using algorithms based on variational methods, message passing, and Markov chain Monte Carlo (MCMC) simulation. Over the years, I’ve taught a dozen courses on machine learning and Bayesian networks to more than a thousand students in all.
In 2005, I became a senior fellow in the neural computation program of the Canadian Institute for Advanced Research, an amazing opportunity to share ideas and collaborate with leaders in the field, such as Yann LeCun, Yoshua Bengio, Yair Weiss, and the director, Geoff Hinton.
DB: What got you started in genomics?
BF: It’s a personal story. In 2002, a couple years into my new role as a professor at the University of Toronto, my wife at the time and I learned that the baby she was carrying had a genetic problem. The counselor we met didn’t do much to clarify things: she could only suggest that either nothing was wrong, or that, on the other hand, something may be terribly wrong. That experience, incredibly difficult for many reasons, also put my professional life into sharp relief: the mainstay of my work, say, in detecting cats in YouTube videos, seemed less significant—all things considered.
I learned two lessons: first, I wanted to use machine learning to improve the lives of hundreds of millions of people facing similar genetic challenges. Second, reducing uncertainty is tremendously valuable: giving someone news, either good or bad, lets them plan accordingly. In contrast, uncertainty is usually very difficult to process.
With that, my research goals changed in kind. Our focus pivoted to understanding how the genome works using deep learning.
DB: Why do you think machine learning plus genome biology is important?
BF: Genome biology, as a field, is generating torrents of data. You will soon be able to sequence your genome using a cell-phone size device for less than a trip to the corner store. And yet, the genome is only part of the story: there exists huge amounts of data that describe cells and tissues. We, as humans, can’t quite grasp all this data: we don’t yet know enough biology. Machine learning can help solve the problem.
At the same time, others in the machine learning community recognize this need. At last year’s premier conference on machine learning, four panelists—Yann LeCun, director of AI at Facebook; Demis Hassabis, co-founder of DeepMind; Neil Lawrence, professor at the University of Sheffield; and Kevin Murphy from Google—identified medicine as the next frontier for deep learning.
To succeed, we need to bridge the “genotype-phenotype divide.” Genomic and phenotype data abound. Unfortunately, the state-of-the-art in meaningfully connecting these data results in a slow, expensive, and inaccurate process of literature searches and detailed wetlab experiments. To close the loop, we need systems that can determine intermediate phenotypes called “molecular phenotypes,” which function as stepping stones from genotype to disease phenotype. For this, machine learning is indispensable.
As we speak, there’s a new generation of young researchers using machine learning to study how genetics impact molecular phenotypes, in groups such as Anshul Kundaje’s at Stanford. To name just a few of these upcoming leaders: Andrew Delong, Babak Alipanahi, and David Kelley of the University of Toronto and Harvard, who study protein-DNA interactions; Jinkuk Kim of MIT, who studies gene repression; and Alex Rosenberg, who is developing experimental methods for examining millions of mutations and their influence on splicing at the University of Washington. In parallel, I think it’s exciting to see an emergence of startups working in this field, such as Atomwise, Grail and others.
DB: What was the state of the genomics field when you started to explore it?
BF: Researchers used a variety of simple “linear” machine learning approaches, such as support vector machines and linear regression that could, for instance, predict cancer from a patient’s gene expression pattern. These techniques were, by their design, “shallow.” In other words, each input to the model would net a very simple “advocate” or “don’t advocate” for the class label. Those methods didn’t account for the complexity of biology.
Hidden Markov models and related techniques for analyzing sequences became popular in the 1990s and early 2000s. Richard Durbin and David Haussler were leading groups in this area.
Around the same time, Chris Burge’s group at MIT developed a Markov model that could detect genes, inferring the beginning of the gene as well as the boundaries between different parts, called introns and exons. These methods were useful for low-level “sequence analysis,” but they did not bridge the genotype-phenotype divide.
Broadly speaking, the state of research at the time was driven by primarily shallow techniques that did not sufficiently account for the underlying biological mechanisms for how the text of the genome gets converted into cells, tissues, and organs.
DB: What does it mean to develop computational models that sufficiently account for the underlying biology?
BF: One of the most popular ways of relating genotype to phenotype is to look for mutations that correlate with disease, in what’s called a genome-wide association study (GWAS). This approach is also shallow in the sense that it discounts the many biological steps involved in going from a mutation to the disease phenotype. GWAS methods can identify regions of DNA that may be important, but most of the mutations they identify aren’t causal. In most cases, if you could “correct” the mutation, it wouldn’t affect the phenotype.
A very different approach accounts for the intermediate molecular phenotypes. Take gene expression, for example. In a living cell, a gene gets expressed when proteins interact in a certain way with the DNA sequence upstream of the gene—i.e., the “promoter.” A computational model that respects biology should incorporate this promoter-to-gene expression chain of causality. In 2004, Beer and Tavazoie wrote what I considered an inspirational paper. They sought to predict every yeast gene’s expression level based on its promoter sequence, using logic circuits that took as input features derived from the promoter sequence. Ultimately, their approach didn’t pan out, but it was a fascinating endeavor nonetheless.
My group’s approach was inspired by Beer and Tavazoie’s work, but differed in three ways: we examined mammalian cells, we used more advanced machine learning techniques, and we focused on splicing instead of transcription. This last difference was a fortuitous turn in retrospect. Transcription is far more difficult to model than splicing. Splicing is a biological process wherein some parts of the gene (introns) are removed and the remaining parts (exons) are connected together. Sometimes exons are removed, too, and this can have a major impact on phenotypes, including neurological disorders and cancers.
To crack splicing regulation using machine learning, my team collaborated with a group led by an excellent experimental biologist named Benjamin Blencowe. We built a framework for extracting biological features from genomic sequences, pre-processing the noisy experimental data, and training machine learning techniques to predict splicing patterns from DNA. This work was quite successful, and led to several publications in “Nature” and “Science.”
DB: Is genomics different from other applications of machine learning?
BF: We discovered that genomics entails unique challenges, compared to vision, speech, and text processing. A lot of the success in vision rests on the assumption that the object to be classified occupies a substantial part of the input image. In genomics, the difficulty emerges because the object of interest occupies only a tiny fraction—say, one millionth—of the input. Put another way, your classifier acts on trace amounts of signal. Everything else is noise—and a lot of it. Worse yet, it’s relatively structured noise comprised of other, much larger objects irrelevant to the classification task. That’s genomics for you.
The more concerning complication is that we don’t ourselves really know how to interpret the genome. When we inspect a typical image, we naturally recognize its objects, and by extension, we know what we want the algorithm to look for. This applies equally well to text analysis and speech processing, domains in which we have some handle on the truth. In stark contrast, humans are not naturally good at interpreting the genome. In fact, they’re very bad at it.
All this is to say that we must turn to truly superhuman artificial intelligence to overcome our limitations.
DB: Can you tell us more about your work around medicine?
BF: We set out to train our systems to predict molecular phenotypes without including any disease data. Yet, once it was trained, we realized our system could in fact make accurate predictions for disease; it learned how the cell reads the DNA sequence and turns it into crucial molecules. Once you have a computational model of how things work normally, you can use it to detect when things go awry.
We then directed our system to large-scale disease mutation data sets. Suppose there is some particular mutation in the DNA. We feed that mutated DNA sequence, as well as its non-mutated counterpart, into our system and compare the two outputs, the molecular phenotypes. If we observe a big change, we label the mutation as potentially pathogenic. It turns out that this approach works well.
But of course, it isn’t perfect. First, the mutation may change the molecular phenotype, but not lead to disease. Second, the mutation may not affect the molecular phenotype that we’re modeling, but lead to a disease in some other way. Third, of course, our system isn’t perfectly accurate. Despite these shortcomings, our approach can accurately differentiate disease from benign mutations. Last year, we published papers in “Science” and “Nature Biotechnology” demonstrating that the approach is significantly more accurate than competing ones.
DB: Where is your company, Deep Genomics, headed?
BF: Our work requires specialized skills from a variety of areas, including deep learning, convolutional neural networks, random forests, GPU computing, genomics, transcriptomics, high-throughput experimental biology, and molecular diagnostics. For instance, we have on board Hui Xiong, who invented a Bayesian deep learning algorithm for predicting splicing, and Daniele Merico, who developed the whole genome sequencing diagnostics system used at the Hospital for Sick Children. We will continue to recruit talented people in these domains.
Broadly speaking, our technology can impact medicine in numerous ways, including: genetic diagnostics, refining drug targets, pharmaceutical development, personalized medicine, better health insurance and even synthetic biology. Right now, we are focused on diagnostics, as it’s a straightforward application of our technology. Our engine provides a rich source of information that can be used to make more reliable patient decisions at lower cost.
Going forward, many emerging technologies in this space will require the ability to understand the inner workings of the genome. Take, for example, gene editing using the CRISPR/Cas9 system. This technique let’s us “write” to DNA and, as such, could be a very big deal down the line. That said, knowing how to write is not the same as knowing what to write. If you edit DNA, it may make the disease worse, not better. Imagine instead if you could use a computational “engine” to determine the consequences of gene editing writ large. That is, to be fair, a ways off. Yet ultimately, that’s what we want to build.
