Chapter 1. Installation and Quick-Start
You will be happy to know that H2O is very easy to install. First I will show how to install it with R, using CRAN, and then how to install it with Python, using pip.1
After that we will dive into our first machine learning project: load some data, make a model, make some predictions, and evaluate success. By that point you will be able to boast to family, friends, and the stranger lucky enough to sit next to you on the bus that you’re a bit of an expert when it comes to deep learning and all that jazz.
After a detour to look at how random elements can lead us astray, the chapter will close with a look at the web interface, Flow, that comes with H2O.
Preparing to Install
The examples in this book are going to be in R and Python. So you need one of those already installed. And you will need Java. If you have the choice, I recommend you use 64-bit versions of everything, including the OS. (In download pages, 64-bit versions are often labeled with “x64,” while 32-bit versions might say “x86.”)
You may wonder if the choice of R or Python matters? No, and why will be explained shortly. There is also no performance advantage to using scripts versus more friendly GUI tools such as Jupyter or RStudio.
Installing R
On Linux your distro’s package manager should make this trivial: sudo apt-get install r-base on Debian/Ubuntu/Mint/etc., and sudo yum install R on RedHat/Fedora/Centos/etc.
Mac users should head to https://cran.r-project.org/bin/macosx/ and follow the instructions.
On Windows go to http://cran.rstudio.com/bin/windows/ and download and run the exe, then follow the prompts. On the Select Components page it wants to install both the 32-bit and 64-bit versions; I chose to only install 64-bit, but there is no harm in installing both.
The optional second step of an R install is to install RStudio; you can do everything from the command line that you need to run H2O, but RStudio makes everything easier to use (especially on Windows, where the command line is still stuck in 1995). Go to https://www.rstudio.com/products/rstudio/download/, download, and install it.
Installing Python
H2O works equally well with Python 2.7 or Python 3.5, as should all the examples in this book. If you are using an earlier version of Python you may need to upgrade. You will also need pip, Python’s package manager.
On Linux, sudo apt-get python-pip on Debian/Ubuntu/Mint/etc.; or for Python 3, it is sudo apt-get python3-pip. (Python is a dependency of pip, so by installing pip we get Python too.) For RedHat/Fedora/Centos/etc., the best command varies by exactly which version you are using, so see the latest Linux Python instructions.
On a Mac, see Using Python on a Macintosh.
On Windows, see Using Python on Windows. Remember to choose a 64-bit install (unless you are stuck with a 32-bit version of Windows, of course).
Tip
You might also want to take a look at Anaconda. It is a Python distribution containing almost all the data science packages you are likely to want. As a bonus, it can be installed as a normal user, which is helpful for when you do not have root access. Linux, Mac, and Windows versions are available.
Privacy
H2O has some code2 to call Google Analytics every time it starts. This appears to be fairly anonymous, and is just for tracking which versions are being used, but if it bothers you, or would break company policy, creating an empty file called .h2o_no_collect in your home directory ("C:\Users\YourName\" on Windows) stops it. You’ll know that works if you see “Opted out of sending usage metrics.” in the info log. Another way to opt out it is given in “Running from the Command Line” in Chapter 10.
Installing Java
You need Java installed, which you can get at the Java download page. Choose the JDK.3 If you think you have the Java JDK already, but are not sure, you could just go ahead and install H2O, and come back and (re-)install Java if you are told there is a problem.
For instance, when testing an install on 64-bit Windows, with 64-bit R, it was when I first tried library(h2o) that I was told I had a 32-bit version of the JDK installed. After a few seconds glaring at the screen, I shrugged, and downloaded the latest version of the JDK. I installed it, tried again, and this time everything was fine.
Install H2O with R (CRAN)
(If you are not using R, you might want to jump ahead to “Install H2O with Python (pip)”.)
Start R, and type install.packages("h2o"). Golly gosh, when I said it was easy to install, I meant it! That command takes care of any dependencies, too.
If this is your first time using CRAN4 it will ask for a mirror to use. Choose one close to you. Alternatively, choose one in a place you’d like to visit, put your shades on, and take a selfie.
If you want H2O installed site-wide (i.e., usable by all users on that machine), run R as root, sudo R, then type install.packages("h2o").
Let’s check that it worked by typing library(h2o). If nothing complains, try the next step: h2o.init(). If the gods are smiling on you then you’ll see lots of output about how it is starting up H2O on your behalf, and then it should tell you all about your cluster, something like in Figure 1-1. If not, the error message should be telling you what dependency is missing, or what the problem is.
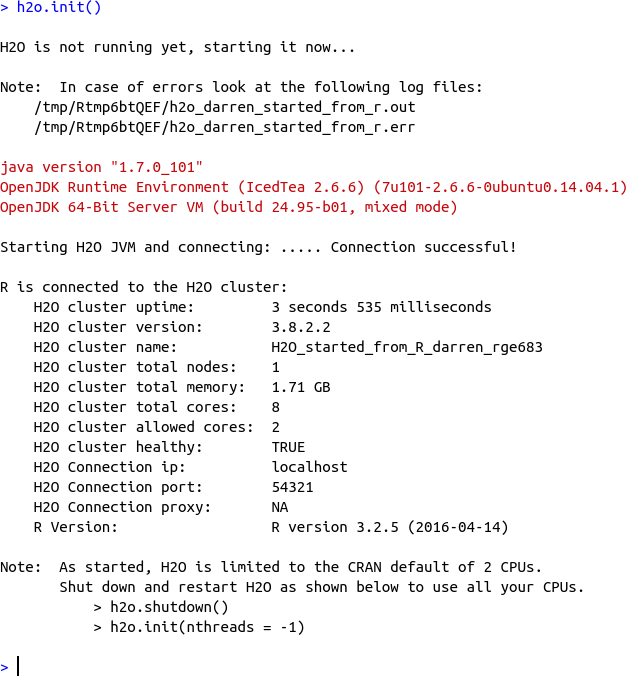
Figure 1-1. Running h2o.init() (in R)
Let’s just review what happened here. It worked. Therefore5 the gods are smiling on you. The gods love you! I think that deserves another selfie: in fact, make it a video of you having a little boogey-woogey dance at your desk, then post it on social media, and mention you are reading this book. And how good it is.
The version of H2O on CRAN might be up to a month or two behind the latest and greatest. Unless you are affected by a bug that you know has been fixed, don’t worry about it.
h2o.init() will only use two cores on your machine and maybe a quarter of your system memory,6 by default. Use h2o.shutdown() to, well, see if you can guess what it does. Then to start it again, but using all your cores: h2o.init(nthreads = -1). And to give it, say, 4GB and all your cores: h2o.init(nthreads = -1, max_mem_size = "4g").
Install H2O with Python (pip)
(If you are not interested in using Python, skip ahead to “Our First Learning”.)
From the command line, type pip install -U h2o. That’s it. Easy-peasy, lemon-squeezy.
The -U just says to also upgrade any dependencies. On Linux you probably needed to be root, so instead type sudo pip install -U h2o. Or install as a local user with pip install -U --user h2o.
To test it, start Python, type import h2o, and if that does not complain, follow it with h2o.init(). Some information will scroll past, ending with a nice table showing, amongst other things, the number of nodes, total memory, and total cores available, something like in Figure 1-2.7 (If you ever need to report a bug, make sure to include all the information from that table.)
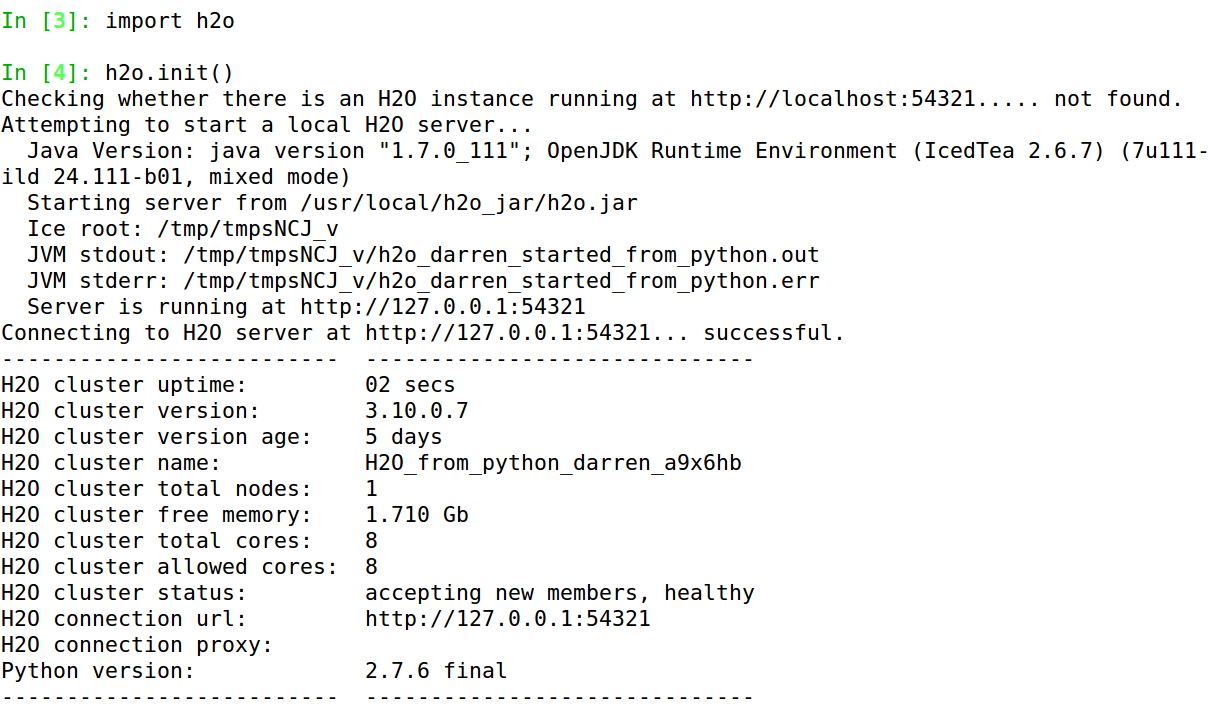
Figure 1-2. Running h2o.init() (in Python)
If you do indeed see that table, stand up and let out a large whoop. Don’t worry about what your coworkers think. They love you and your eccentricities. Trust me.
By default, your H2O instance will be allowed to use all your cores, and (typically) 25% of your system memory. That is often fine but, for the sake of argument, what if you wanted to give it exactly 4GB of your memory, but only two of your eight cores? First shut down H2O with h2o.shutdown(), then type h2o.init(nthreads=2, max_mem_size=4). The following excerpt from the information table confirms that it worked:
... H2O cluster total free memory: 3.56 GB H2O cluster total cores: 8 H2O cluster allowed cores: 2 ...
Note
Using virtualenv does not work with H2O.8 To be precise, it installs but cannot start H2O for you. If you really want to install it this way, follow the instructions on starting H2O from the command line in Chapter 10. The h2o.init(), and everything else, will then work.
Our First Learning
Now that we have everything installed, let’s get down to business. The Python and R APIs are so similar that we will look at them side-by-side for this example. If you are using Python look at Example 1-1, and if you are using R take a look at Example 1-2. They repeat the import/library and h2o.init code we ran earlier; don’t worry, this does no harm.
I’m going to spend a few pages going through this in detail, but I want to just emphasize that this is the complete script: it downloads data, prepares it, creates a multi-layer neural net model (i.e., deep learning) that is competitive with the state of the art on this data set, and runs predictions on it.
Example 1-1. Deep learning on the Iris data set, in Python
importh2oh2o.init()datasets="https://raw.githubusercontent.com/DarrenCook/h2o/bk/datasets/"data=h2o.import_file(datasets+"iris_wheader.csv")y="class"x=data.namesx.remove(y)train,test=data.split_frame([0.8])m=h2o.estimators.deeplearning.H2ODeepLearningEstimator()m.train(x,y,train)p=m.predict(test)
Example 1-2. Deep learning on the Iris dataset, in R
library(h2o)h2o.init(nthreads=-1)datasets<-"https://raw.githubusercontent.com/DarrenCook/h2o/bk/datasets/"data<-h2o.importFile(paste0(datasets,"iris_wheader.csv"))y<-"class"x<-setdiff(names(data),y)parts<-h2o.splitFrame(data,0.8)train<-parts[[1]]test<-parts[[2]]m<-h2o.deeplearning(x,y,train)p<-h2o.predict(m,test)
![]() ,
, ![]() ,
, ![]() are preparing the data,
are preparing the data, ![]() is training the model, and
is training the model, and ![]() is using that model.
is using that model.
![]() illustrates the first major concept we need to understand when using H2O: all the data is on the cluster (the server), not on our client. Even when client and cluster are the same machine.
illustrates the first major concept we need to understand when using H2O: all the data is on the cluster (the server), not on our client. Even when client and cluster are the same machine.
Therefore, whenever we want to train a model, or make a prediction, we have to get the data into the H2O cluster; we will look at that topic in more depth in Chapter 2. For now, just appreciate that this line has created a frame on the cluster called “iris_wheader.hex.” It recognized the first line of the csv file was a header row, so it has automatically named the columns. It has also realized (from analyzing the data) that the “class” column was categorical, which means we will be doing a multinomial categorization, not a regression (see “Jargon and Conventions”).
![]() defines a couple of helper variables:
defines a couple of helper variables: y to be the name of the field we want to learn, and x to be the names of the fields we want to learn from; in this case that means all the other fields. In other words, we will attempt to use the four measurements, sepal_len, sepal_wid, petal_len, and petal_wid, to predict which species a flower belongs to.
![]() (splitting into training and test data) is another big concept, which boils down to trying not to overfit. Briefly, what we are doing is (randomly) choosing 80% of our data to train on, and then we will try using our model on the remaining 20%, to see how well it did. In a production system, this 20% represents the gardeners coming in with new flowers and asking us what species they are.
(splitting into training and test data) is another big concept, which boils down to trying not to overfit. Briefly, what we are doing is (randomly) choosing 80% of our data to train on, and then we will try using our model on the remaining 20%, to see how well it did. In a production system, this 20% represents the gardeners coming in with new flowers and asking us what species they are.
A reminder that the Python code to split the data looked like the following. split_frame() is one of the member functions of class H2OFrame. The [0.8] tells it to put 80% in the first split, the rest in the second split:
train,test=data.split_frame([0.8])
In R, h2o.splitFrame() takes an H2O frame and returns a list of the splits, which are assigned to train and test, for readability:
parts<-h2o.splitFrame(data,0.8)train<-parts[[1]]test<-parts[[2]]
The split, being decided randomly for each row, is roughly 120/30 rows, but you may get a few more training rows, or a few more test rows.12
Let’s quickly recap what we have. As shown in Figure 1-3, the client just has handles (pointers) to the actual data on the H2O cluster.
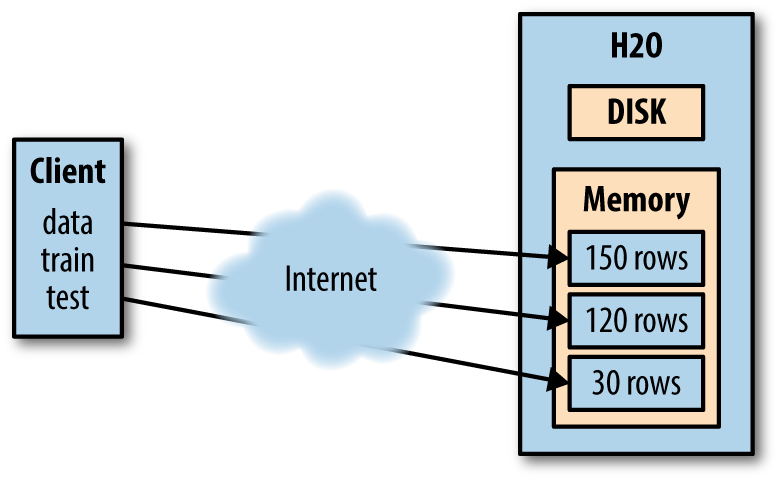
Figure 1-3. Recap of what data is where
Of course, our “cluster” is on localhost, on the same machine as our client, so it is all the same system memory. But you should be thinking as if they are on opposite sides of the globe. Also think about how it might be a billion rows, too many to fit in our client’s memory. By adding machines to a cluster, as long as the total memory of the cluster is big enough, it can be loaded, and you can analyze those billion rows from a client running on some low-end notebook.
Training and Predictions, with Python
At last we get to ![]() , the machine learning. In Python it is a two-step process:
, the machine learning. In Python it is a two-step process:
-
Create an object for your machine-learning algorithm, and optionally specify parameters for it:
m=h2o.estimators.deeplearning.H2ODeepLearningEstimator() -
Tell it to train and which data sets to use:
m.train(x,y,train)
If you prefer scikit-learn style, you can instead write:
fromh2o.estimators.deeplearningimportH2ODeepLearningEstimatorm=H2ODeepLearningEstimator()m.train(x,y,train)
No parameters to the constructor means the model is built with all defaults, which means (amongst other things): two hidden layers, each with 200 neurons, and 10 epochs of training. (Chapter 8 will define “neurons” and “epochs"―but don’t go there yet. The important point is that default settings are usually quite quick to train, just a few seconds on this iris data.)
As with the data frames, m is a class wrapper around a handle, pointing to the actual model stored on the H2O cluster. If you print m you get a lot of details of how the training went, or you can use member functions to pull out just the parts you are interested in—e.g., m.mse() tells me the MSE (mean squared error) is 0.01097. (There is a random element, so you are likely to see slightly different numbers.13)
m.confusion_matrix(train) gives the confusion matrix, which not only shows how many in each category it got right, but which category is being chosen when it got them wrong. The results shown here are on the 120 training samples:
| Iris-setosa | Iris-versicolor | Iris-virginica | Error | Rate |
|---|---|---|---|---|
42 |
0 |
0 |
0 |
0 / 42 |
0 |
37 |
1 |
0.0263158 |
1 / 38 |
0 |
1 |
39 |
0.025 |
1 / 40 |
42 |
38 |
40 |
0.0166667 |
2 / 120 |
In this case I see it matched all 42 setosa perfectly, but it thought one versicolor was a virginica, and one virginica was a versicolor.
The final line of the listing, ![]() , was
, was p = m.predict(test), and it makes predictions using this model, and puts them in p. Here are a few of the predictions. The leftmost column shows which category it chose. The other columns show the probability it has assigned for each category for each test sample. You can see it is over 99.5% certain about all its answers here:
| predict | Iris-setosa | Iris-versicolor | Iris-virginica |
|---|---|---|---|
Iris-setosa |
0.999016 |
0.000983921 |
1.90283E-019 |
Iris-setosa |
0.998988 |
0.00101178 |
1.40209E-020 |
Iris-versicolor |
5.22035E-005 |
0.997722 |
0.00222536 |
Iris-versicolor |
0.000275126 |
0.995354 |
0.00437055 |
Just as before, this is a frame on the H2O cluster, so when you see it, you only you see a preview, the first 10 rows. To see all 30 predictions you need to download, which is done with p.as_data_frame(). If you don’t have pandas installed, you get a nested list, something like this:
[['predict', 'Iris-setosa', 'Iris-versicolor', 'Iris-virginica'], ['Iris-setosa', '0.9990160791818314', '9.83920818168421E-4', '1.9028267028039464E-19'], ['Iris-setosa', '0.9989882189908829', ... ..., ['Iris-virginica', '1.72617432126E-11', '1.0197263306598747E-4', '0.9998980273496721']]
You could do analysis with that. However, H2O’s Python API integrates with pandas, and if you are using Python for data work, chances are you already know and use it. (If not, pip install pandas should be all you need to install it.) As long as you have installed pandas, p.as_data_frame() will instead give:
predict Iris-setosa Iris-versicolor Iris-virginica 0 Iris-setosa 9.990161e-01 0.000984 1.902827e-19 1 Iris-setosa 9.989882e-01 0.001012 1.402089e-20 ...
What else can we do? Chapter 2 will delve into this topic further, but how about (p["predict"] == test["class"]).mean() to tell us the percentage of correct answers? Or p["predict"].cbind(test["class"]).as_data_frame() to give a two-column output of each prediction against the correct answer:
predict class 0 Iris-setosa Iris-setosa 1 Iris-setosa Iris-setosa ... 11 Iris-versicolor Iris-versicolor 12 Iris-virginica Iris-versicolor 13 Iris-versicolor Iris-versicolor 14 Iris-virginica Iris-versicolor 15 Iris-versicolor Iris-versicolor ... 28 Iris-virginica Iris-virginica 29 Iris-virginica Iris-virginica
Training and Predictions, with R
In R, ![]() , the machine learning is a single function call, with parameters and training data being given at the same time. As a reminder, the command was:
, the machine learning is a single function call, with parameters and training data being given at the same time. As a reminder, the command was: m <- h2o.deeplearning(x, y, train). (In fact, I used m <- h2o.deeplearning(x, y, train, seed = 99, reproducible = TRUE) to get repeatable results, but you generally don’t want to do that as it will only use one core and take longer.)
Just like with the data, the model is stored on the H2O cluster, and m is just a handle to it. h2o.mse(m) tells me the mean squared error (MSE) was 0.01097. h2o.confusionMatrix(m) gives the following confusion matrix (on the training data, by default):
Confusion Matrix: vertical: actual; across: predicted
setosa versicolor virginica Error Rate
setosa 42 0 0 0.0000 = 0 / 42
versicolor 0 37 1 0.0263 = 1 / 38
virginica 0 1 39 0.0250 = 1 / 40
Totals 42 38 40 0.0167 = 2 / 120
So, a perfect score on the setosa, but one versicolor wrong—it thought it was a virginica—and one virginica it thought was a versicolor. The bottom right tells us it therefore had an error rate of 1.67%. (Remember this was on the data it had seen.)
The final line of the listing, ![]() , was
, was p <- h2o.predict(m, test) and it makes predictions using the model m. Again, p is a handle to a frame on the H2O server. If I output p I only see the first six predictions. To see all of them I need to download the data. When working with remote clusters, or big data… sorry, Big Data™, be careful here: you will first want to consider how much of your data you actually need locally, how long it will take to download, and if it will even fit on your machine.
By typing as.data.frame(p) I see all 30 predictions (just a few shown here):
predict Iris-setosa Iris-versicolor Iris-virginica ----------- ----------- --------------- -------------- Iris-setosa 0.999016 0.0009839 1.90283e-19 Iris-setosa 0.998988 0.0010118 1.40209e-20 Iris-setosa 0.999254 0.0007460 9.22466e-19 ... Iris-virginica 1.5678e-08 0.3198963 0.680104 Iris-versicolor 2.3895e-08 0.9863869 0.013613 ... Iris-virginica 3.9084e-14 2.192105e-06 0.999998
The predict column in the first row is the class it is predicting for the first row in the test data. The other three columns show its confidence. You can see it is really sure that it was a setosa. If you explore the predictions you will see it is less sure of some of the others.
The next question you are likely to have is which ones, if any, did H2O’s model get wrong? The correct species is in test$class, while deep learning’s guess is in p$predict. There are two approaches so, based on what you know so far, have a think about the difference between this:
as.data.frame(h2o.cbind(p$predict,test$class))
and:
cbind(as.data.frame(p$predict),as.data.frame(test$class))
In the first approach, p$predict and test$class are combined in the cluster to make a new data frame in the cluster. Then this new two-column data frame is downloaded. In the second approach, one column from p is downloaded to R, then one column from test is downloaded, and then they are combined in R’s memory, to make a two-column data frame. As a rule of thumb, prefer the first way.
In my case (your results might differ slightly) this gives (I’ve put an asterisk by the two cases it got wrong):
predict class 1 setosa setosa 2 setosa setosa 3 setosa setosa 4 setosa setosa 5 setosa setosa 6 setosa setosa 7 setosa setosa 8 setosa setosa 9 versicolor versicolor 10 versicolor versicolor 11 versicolor versicolor 12 versicolor versicolor 13 virginica versicolor * 14 versicolor versicolor 15 virginica versicolor * 16 versicolor versicolor 17 versicolor versicolor 18 versicolor versicolor 19 versicolor versicolor 20 versicolor versicolor 21 virginica virginica 22 virginica virginica 23 virginica virginica 24 virginica virginica 25 virginica virginica 26 virginica virginica 27 virginica virginica 28 virginica virginica 29 virginica virginica 30 virginica virginica
Another way we could analyze our results is by asking what percentage the H2O model got right. In R that can be done with mean(p$predict == test$class), which tells me 0.933. In other words, the model guessed 93.3% of our unseen 30 test samples correctly, and got 6.7% wrong. As we will see in “On Being Unlucky”, you almost certainly got 0.900 (3 wrong), 0.933 (2 wrong), 0.967 (1 wrong), or 1.000 (perfect score).
Performance Versus Predictions
There is another way we could have found out what percentage it got right. It is to not use predict() at all but instead use h2o.performance(m, test) in R, or m.model_performance(test) in Python. This doesn’t tell us what the individual predictions were, but instead gives us lots of statistics:
ModelMetricsMultinomial: deeplearning ** Reported on test data. ** MSE: 0.0390774346788 R^2: 0.934384904457 LogLoss: 0.122561507096 Confusion Matrix: vertical: actual; across: predicted Iris-setosa Iris-versicolor Iris-virginica Error Rate ------------- ----------------- ---------------- --------- ------ 8 0 0 0 0 / 8 0 10 2 0.166667 2 / 12 0 0 10 0 0 / 10 8 10 12 0.0666667 2 / 30 Top-3 Hit Ratios: k hit_ratio --- ----------- 1 0.933333 2 1 3 1
The hit ratio section at the end tells us the same 0.933 number. (The 1 in the second row means it was 100% accurate if allowed two guesses.) Above that, the confusion matrix tells us that it incorrectly guessed two virginica samples as being versicolor.
Warning
If we study the confidence of our predictions we see the correct answers are mostly over 0.99, with the least-confident correct answer being 0.97. What about our incorrect answers? Test row 13 was 0.45 versus 0.55 (the machine-learning version of a teenager’s sullen shrug) and test row 15 was 0.07 versus 0.93.
This is great, as it means we can mark results with confidence below 0.97 as suspicious. In a medical application that could mean doing another test to get a second opinion; in a financial trading application (or a poker app) it could mean sit this one out, too risky.
But!! Hopefully you got suspicious as soon as I said, “This is great!” We’re choosing our cutoff criteria of 0.97 based on looking at our test results, after being told the correct answers. All parameters used to interpret the test results must only be based on our training data. (“Valid Versus Test?” in Chapter 2 will touch on how you could use a validation data set for this, though.)
On Being Unlucky
This is a good time to consider how randomness affects the results. To find out, I tried remaking the model 100 times (using random seeds 1 to 100). 52 times the model got two wrong, and 48 times it got one wrong. Depending on your perspective, the random effect is either minor (93% versus 97%), or half the time it is twice as bad. The result set I analyzed in the previous section was one of the unlucky ones.
What about the way we randomly split the data into training and test data? How did that affect things? I tried 25 different random splits, which ended up ranging from 111/39 to 130/20, and made 20 models on each. (Making these 500 models took about 20 minutes on my computer; sadly this experiment is not so practical with the larger data sets we will use later in the book.)
It seems the randomness in our split perhaps matters more than the randomness in our model,14 because one split gave a perfect score for all of its 20 models (it had 129 rows to train from, 21 to test on), whereas another only averaged 90% (it had 114 to train from, 36 to test on). You are thinking “Aha! The more training data, the better?” Yet the split that had 130 training rows only managed 90% on almost all its models.
But wait, there’s more! The single most important learning from this little experiment, for me at least, was that 85 of the 500 models (17%) gave a perfect score. Typically you will use just one split, and make one model; 17% of the time you’d be tricked into thinking your model parameters were good enough for perfection.
A year or two ago, it was in the news that 64% of psychology experiments (published in top journals) could not be reproduced. I suspect this kind of bad luck15 was involved in a few of them.
Flow
Flow is the name of the web interface that is part of H2O (no extra installation step needed). It is actually just another client, written in CoffeeScript (a JavaScript-like language) this time, making the same web service calls to the H2O backend that the R or Python clients are making. It is fully featured, by which I mean that you can do all of the following:
-
View data you have uploaded through your client
-
Upload data directly
-
View models you have created through your client (and those currently being created!)
-
Create models directly
-
View predictions you have generated through your client
-
Run predictions directly
You can find it by pointing your browser to http://127.0.0.1:54321. Of course, if you started H2O on a nonstandard port, change the :54321 bit, and if you are accessing a remote H2O cluster, change the 127.0.0.1 bit to the server name of any node in the cluster (the public DNS name or IP address, not the private one, if it is a server with both). When you first load Flow you will see the Flow menu, as shown in Figure 1-4.
Data
Let’s import the same Iris data set we did in the R and Python examples. From the start screen click the “importFiles” link, or from the menu at the top of the screen choose Data then Import Files. Paste the location of the csv file into the search box, then select it, then finally click the Import button; see Figure 1-5.
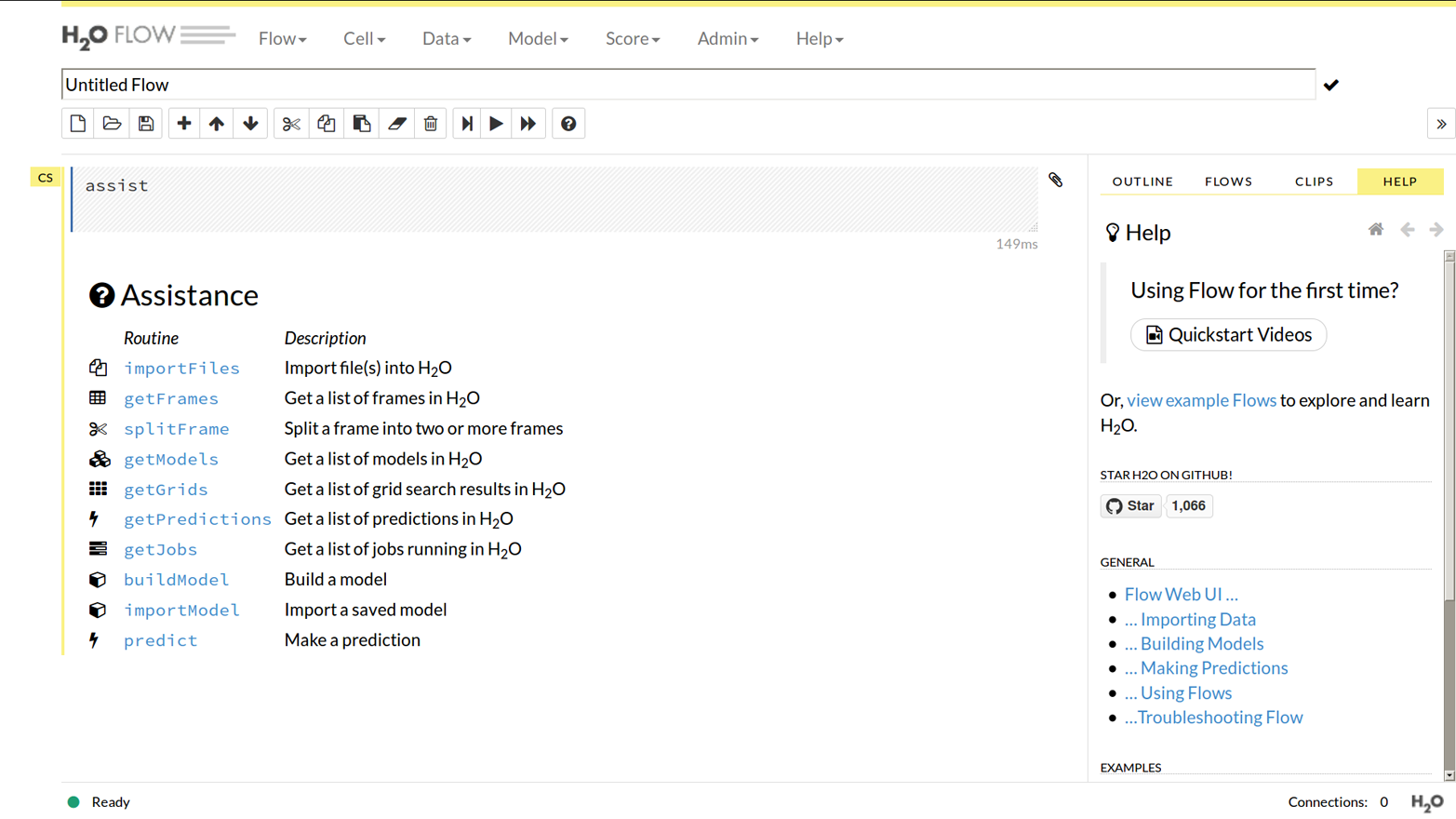
Figure 1-4. The Flow menu
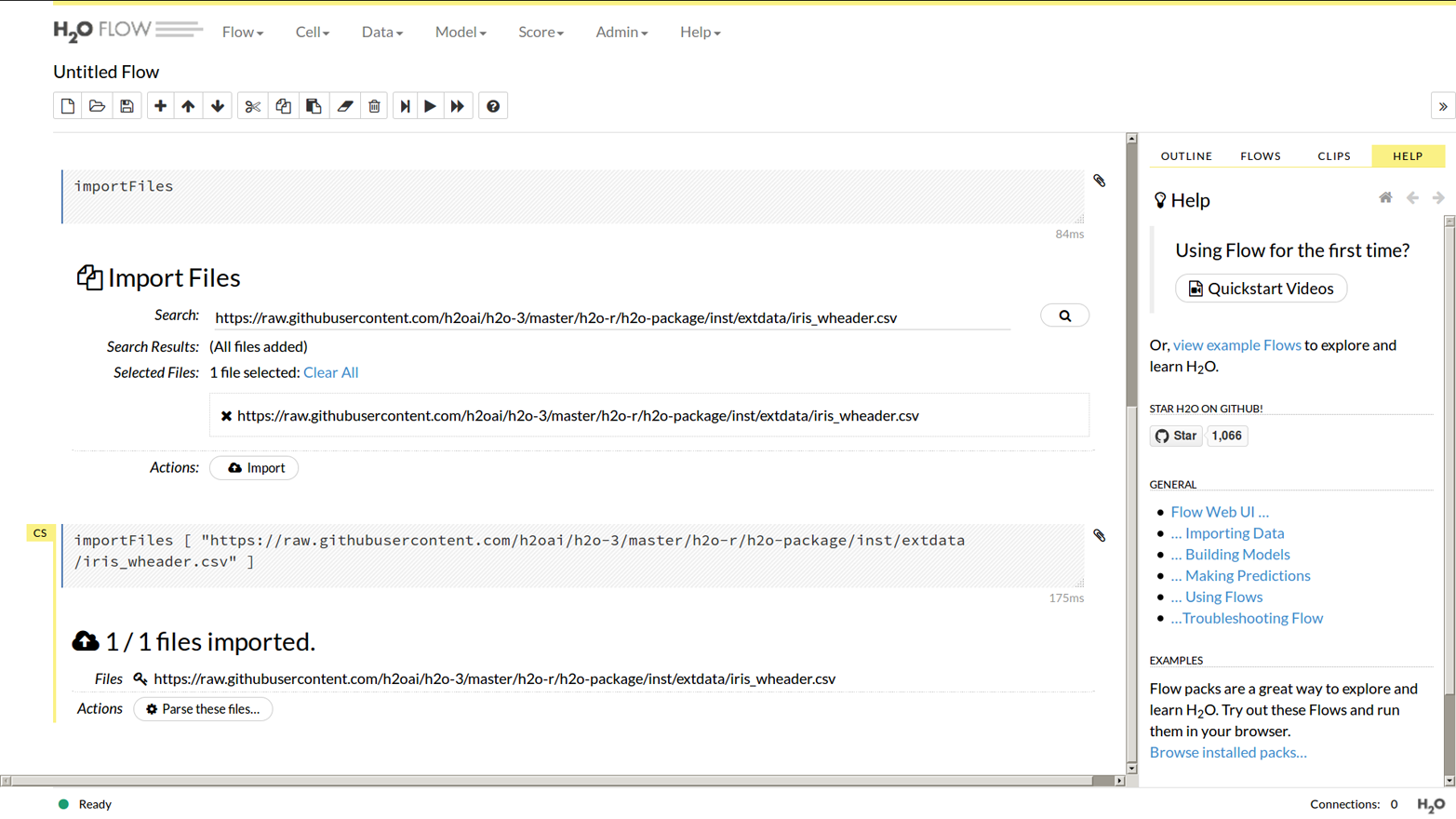
Figure 1-5. Import files
Now click “Parse These Files,” and it gives you the chance to customize the settings as shown in Figure 1-6, but in this case just accepting the defaults is fine.
If you choose “getFrames” from the main menu, either after doing the preceding steps or after loading the data from R or Python, you would see an entry saying “iris_wheader.hex” and that it has 150 rows and 5 columns. If you clicked the “iris_wheader.hex” link you would see Figure 1-7.
You should see there are buttons to split the data (into training/test frames), or build a model, and also that it lists each column. Importantly it has recognized the “class” column as being of type enum, meaning we are ready to do a classification. (If we wanted to do a regression we could click “Convert to numeric” in the Actions column.)
Click Split (the scissors icon), then change the 0.25 to 0.2. Under “Key,” rename the 0.80 split to “train” and the other to “test.”
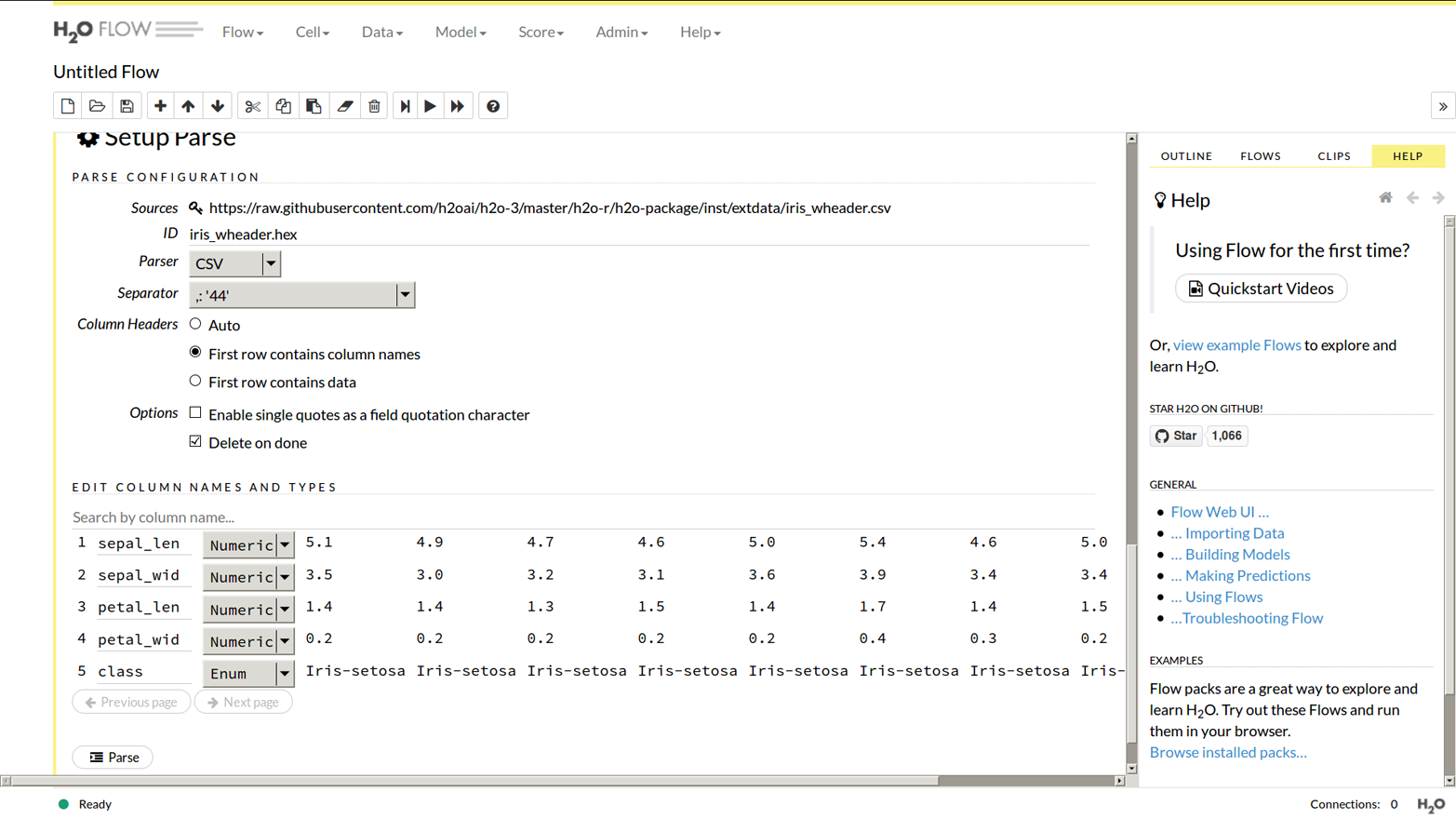
Figure 1-6. Set up file parsing in Flow
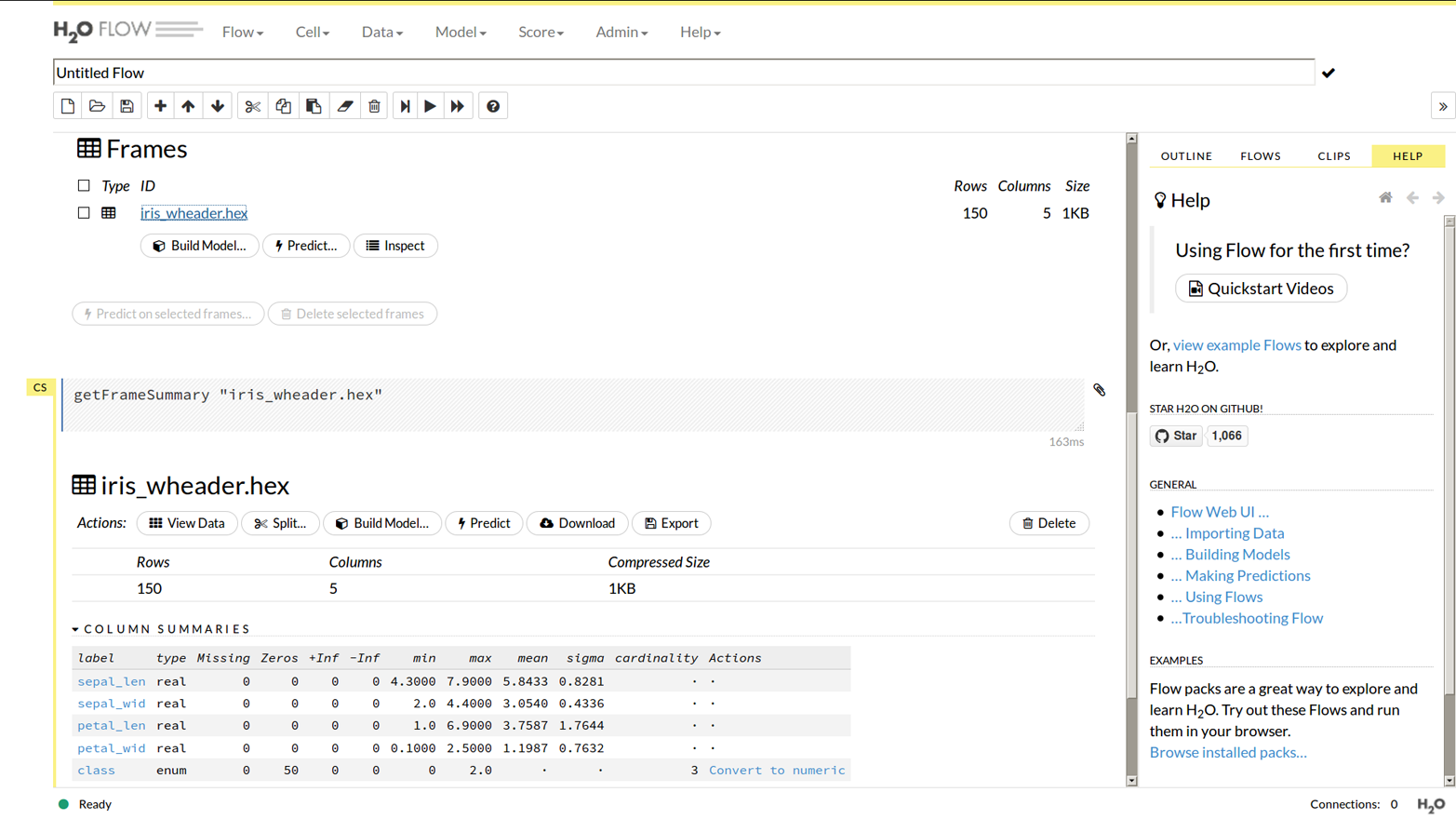
Figure 1-7. Data frame view in Flow
Models
Following on from the previous example, click “train,” then click “Build Model” (a cube icon). From the algorithms, choose Deep Learning.
Loads and loads of parameters appear. You only need to set one of them, near the top: from the “response_column” drop-down, choose “class.” The defaults for everything else are good, so scroll down past them all, and click “Build Model.” You should see something like the output in Figure 1-8.
Now click the “View” button (a magnifying glass icon). Alternatively, if you previously made some models (whether in R, Python, or Flow), choose “Model” from the main menu, then “List All Models,” and click the one of interest. As you can see in Figure 1-9, you get a graphical output; other options allow you to see the parameters the model was built with, or how training progressed.
Predictions
You can do the full load-model-predict cycle in Flow. From the model view click “Predict” (the lightning icon). (Or, choose “Score” from the main menu, then “Predict,” and choose the model from there.)
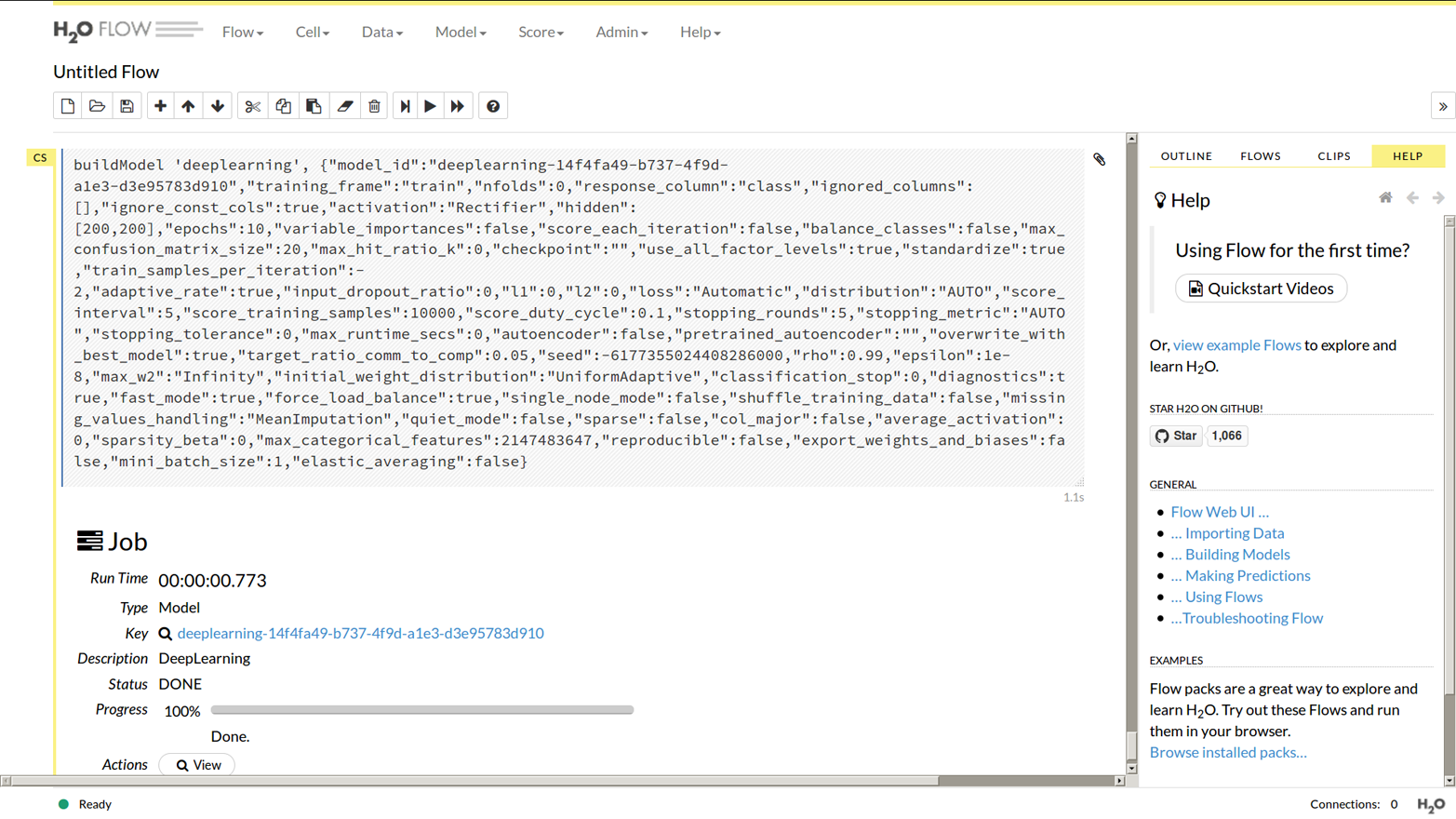
Figure 1-8. A deep learning model in Flow
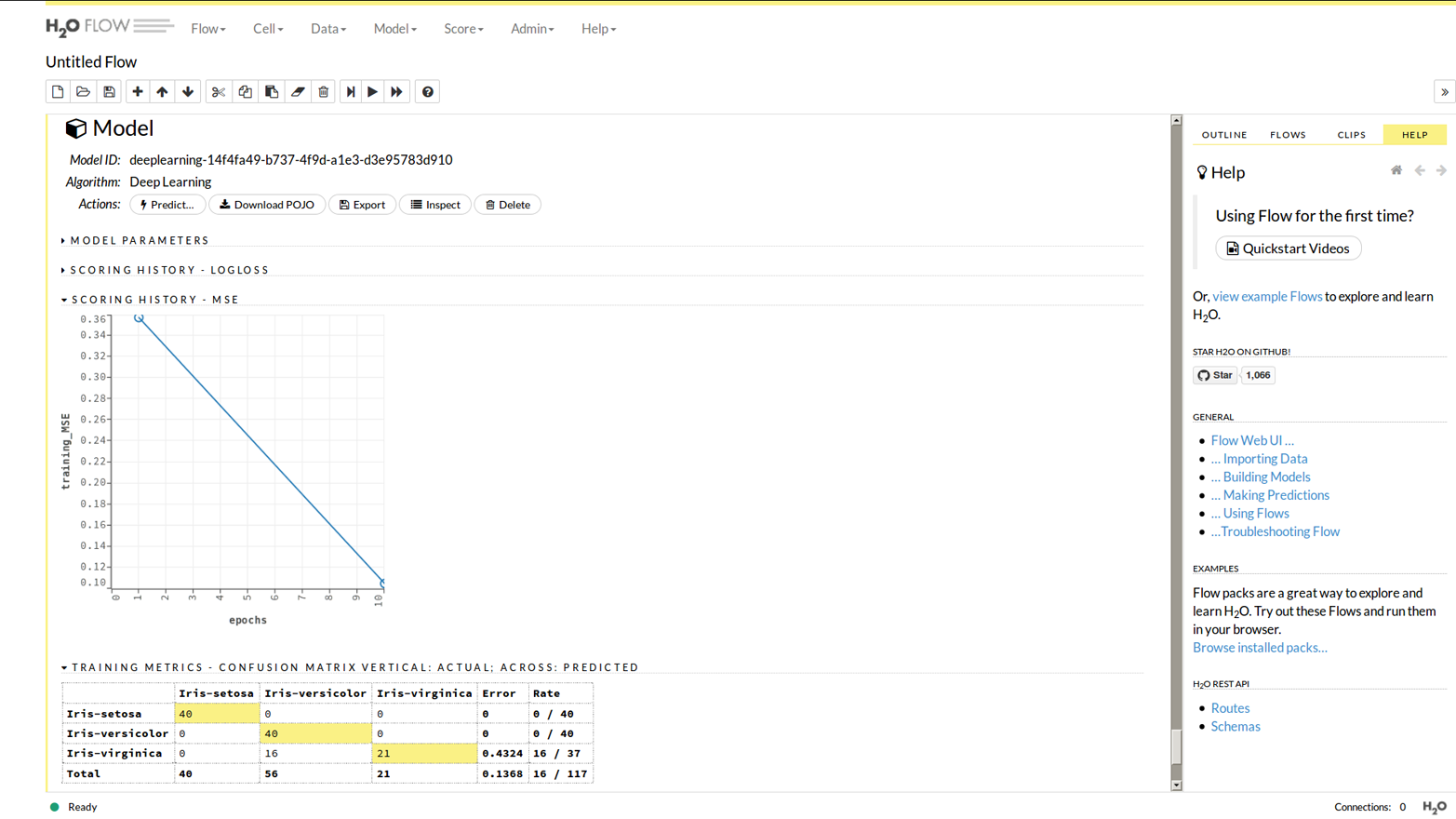
Figure 1-9. Study of a model from Flow
Choose the “test” data frame, and click the Predict button to set it going. You will see results like Figure 1-10.
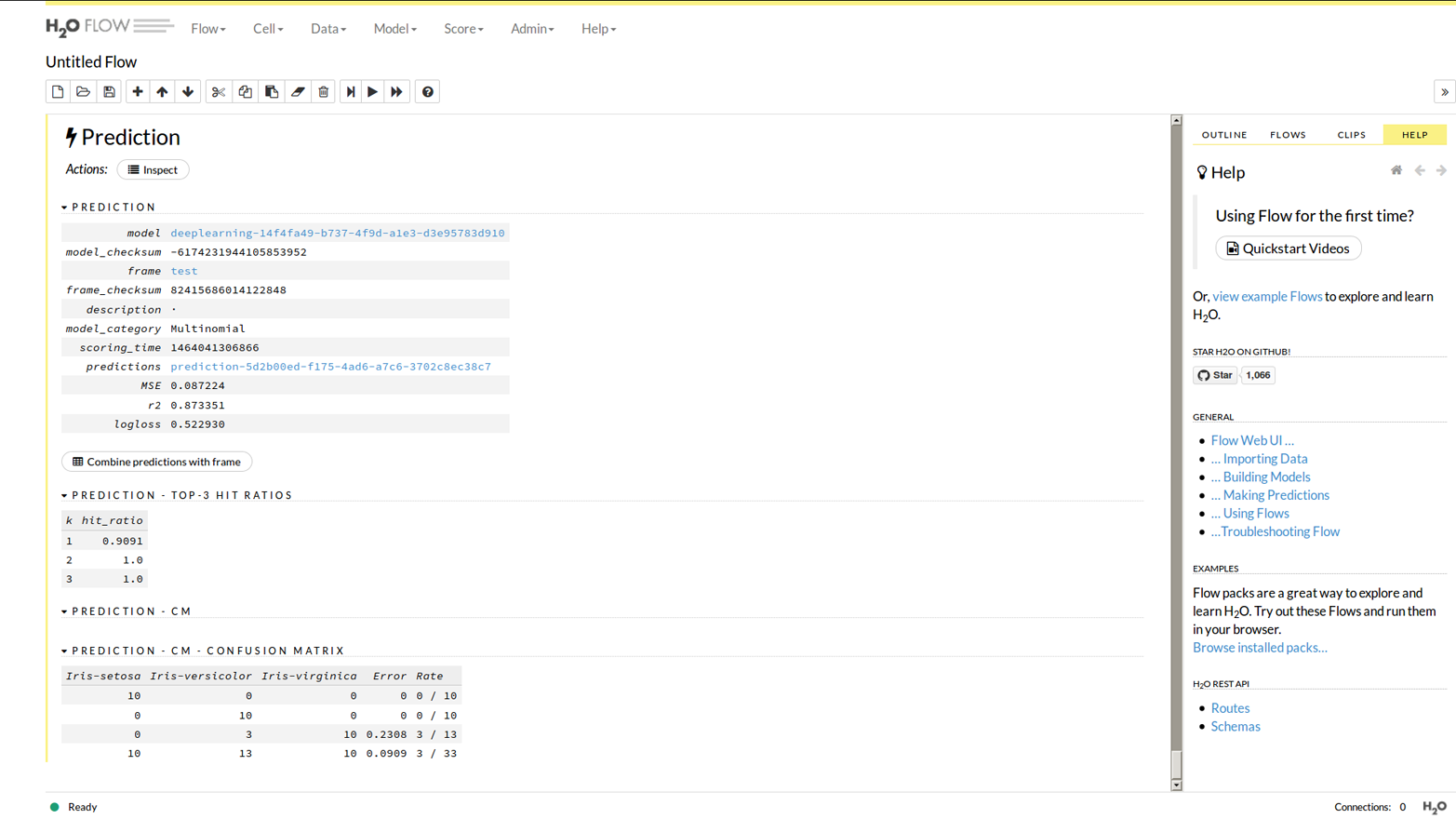
Figure 1-10. A prediction in Flow
Other Things in Flow
The Flow commands you see can be saved as scripts, and loaded back in later. But, there are some things you can do with the R and Python APIs that you cannot do in Flow, principally, merging data sets (either by columns or by rows), and data manipulation (which we will be taking a look at very soon, in Chapter 2).
So, for some users, Flow can do all you need, but most of us will want to use R or Python. I will not be showing Flow examples in the rest of the book, though the knowledge learned as we look at the algorithm parameters in later chapters can be directly applied to models built in Flow.
Having said that, Flow can be useful to you even if you intend to only use R or Python. If you load data from R/Python you can see it in Flow. If you load data in Flow, you can see it in R/Python. Even better, you can start a long-running model from Python or R, then go over to Flow and get immediate feedback on how the training is going. Seeing unexpected performance, you might realize you forgot something, kill it, and thus avoid wasting hours of CPU time. And the Water Meter found under the Admin menu is a very useful way to see how hard each CPU core in your cluster is working.
Summary
In this first chapter we have covered a lot of ground:
-
Installing H2O for R and Python
-
Importing data, making models, and making predictions…
-
…in any of R, Python, or the browser-based Flow UI
And we had a bit of fun, wearing the shades, and deep learning our data like a boss.
We also glossed over quite a few options, and the next chapter (well, in fact, the whole rest of the book) will start digging in deeper. But keep the shades handy, there is lots of fun still to be had.
1 Chapter 10 shows some alternative ways to install H2O (see “Installing the Latest Version”) including how to compile it from source. You might want to do this if you hit a bug that has only been fixed in the latest development version, or if you want to start hacking on H2O itself.
2 http://bit.ly/2f96Hyu as of June 2016.
3 The “Server JRE” or “JRE” choices may work with H2O, but I recommend you always install the JDK.
4 CRAN is R’s package manager. See https://cran.r-project.org/ to learn more about it.
5 It is the ability to make inferences like this that separate the common herd from the data scientist.
6 See http://bit.ly/2gn5h6e for how to query your Java installation and get the default for your system.
7 You perhaps see some deprecation warnings? For the screenshot I ignored them with import warnings;warnings.filterwarnings("ignore", category=DeprecationWarning), but normally I ignore them by turning a blind eye.
8 At least, as of version 3.10.x.x and earlier.
9 The R and Python APIs also call it y; in the REST API it is called response_column.
10 The APIs also call this x, though if you ever poke your nose into the REST API, you will see it actually receives the complement of x: the list of field names to not use, called ignored_columns.
11 If you are unfamiliar with these, the difference between validation and test data, and validation and cross-validation, is covered in Chapter 3.
12 Experiment with seed to get an exact split. For example, h2o.splitFrame(data, 0.8, seed=99) works for me. In Python: data.split_frame([0.8],seed=99).
13 I used h2o.deeplearning(x, y, train, seed = 99, reproducible = TRUE) to get repeatable numbers for this book.
14 This is more common with small data sets like Iris; with larger data sets it is less likely to happen unless, for instance, one category in an enum column is much rarer than other categories.
15 Bad luck from the point of view of a healthier, happier, human society. Of course, it was good luck for the person needing a paper accepted by a journal!
Get Practical Machine Learning with H2O now with the O’Reilly learning platform.
O’Reilly members experience books, live events, courses curated by job role, and more from O’Reilly and nearly 200 top publishers.

