1.4 Reconstruction of Biological Networks
In this section, we describe some existing approaches to reconstruct directed and undirected biological networks from gene expression data and gene sets. To reconstruct directed networks from gene expression data, we present Boolean network, probabilistic Boolean network, and Bayesian network models. We discuss cGraph, frequency method and NICO approaches for network reconstruction using gene sets (Fig 1.4). Next, we present relevance networks and graphical Gaussian models for the reconstruction of undirected biological networks from gene expression data (Fig 1.5). The review of models in case of directed and undirected networks is largely based on Refs. [6–8,17–20] and [2,3,13,32], respectively.
Figure 1.4 (a) Representation of inputs and Boolean data in the frequency method from Ref. [18]. (b) Network inference from PAK pathway [67] using NICO, in the presence of a prior known end points in each path [68]. (c) The building block of cGraph from Ref. [17].
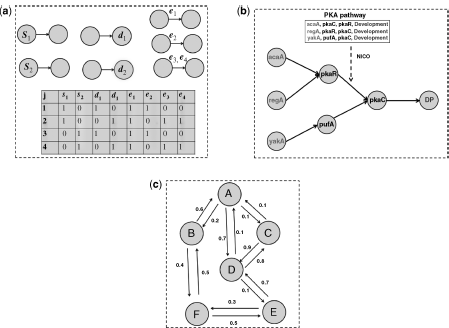
Figure 1.5 Comparison of correlation-based relevance networks (a) and partial correlation based graphical Gaussian modeling (b) performed on a synthetic data set generated from multivariate normal distribution. The figures represent estimated correlations and partial correlations between every pair of genes. Light to dark colors correspond to high to low correlations and partial correlations. ...
Get Statistical and Machine Learning Approaches for Network Analysis now with the O’Reilly learning platform.
O’Reilly members experience books, live events, courses curated by job role, and more from O’Reilly and nearly 200 top publishers.

