Chapter 1. Using R
Machine learning exists at the intersection of traditional mathematics and statistics with software engineering and computer science. In this book, we will describe several tools from traditional statistics that allow you to make sense of that world. Statistics has almost always been concerned with learning something interpretable from data, while machine learning has been concerned with turning data into something practical and usable. This contrast makes it easier to understand the term machine learning: Machine learning is concerned with teaching computers something about the world, so that they can use that knowledge to perform other tasks, while statistics is more concerned with developing tools for teaching humans something about the world, so that they can think more clearly about the world in order to make better decisions.
In machine learning, the learning occurs by extracting as much information from the data as possible (or reasonable) through algorithms that parse the basic structure of the data and distinguish the signal from the noise. After they have found the signal, or pattern, the algorithms simply decide that everything else that’s left over is noise. For that reason, machine learning techniques are also referred to as pattern recognition algorithms. We can “train” our machines to learn about how data is generated in a given context, which allows us to use these algorithms to automate many useful tasks. This is where the term training set comes from, referring to the set of data used to build a machine learning process. The notion of observing data, learning from it, and then automating some process of recognition is at the heart of machine learning, and forms the primary arc of this book.
In this book, we will assume a relatively high degree of knowledge in basic programming techniques and algorithmic paradigms. That said, R remains a relatively niche language even among experienced programmers. In an effort to start everyone at the same starting point, this chapter will also provide some basic information on how to get started using the R language. Later in the chapter we will work through a specific example of using the R language to perform common tasks associated with machine learning.
Warning
This chapter does not provide a complete introduction to the R programming language. As you might expect, no such introduction could fit into a single book chapter. Instead, this chapter is meant to prepare the reader for the tasks associated with doing machine learning in R. Specifically, we describe the process of loading, exploring, cleaning, and analyzing data. There are many excellent resources on R that discuss language fundamentals; such as data types, arithmetic concepts, and coding best practices. Insofar as those topics are relevant to the case studies presented here, we will touch on all of these issues; however, there will be no explicit discussion of these topics. Some of these resources are listed in Table 1-1.
If you have never seen the language and its syntax before, we highly recommend going through this introduction to get some exposure. Unlike other high-level scripting languages, such as Python or Ruby, R has a unique and somewhat prickly syntax and tends to have a steeper learning curve than other languages. If you have used R before, but not in the context of machine learning, there is still value in taking the time to go through this review before moving onto the cases.
R for Machine Learning
R is a language and environment for statistical computing and graphics...R provides a wide variety of statistical (linear and nonlinear modeling, classical statistical tests, time-series analysis, classification, clustering, ...) and graphical techniques, and is highly extensible. The S language is often the vehicle of choice for research in statistical methodology, and R provides an Open Source route to participation in that activity.
—The R Project for Statistical Computing, http://www.r-project.org/
The best thing about R is that it was developed by statisticians. The worst thing about R is that...it was developed by statisticians.
—Bo Cowgill, Google, Inc.
R is an extremely powerful language for manipulating and analyzing data. Its meteoric rise in popularity within the data science and machine learning communities has made it the de facto lingua franca for analytics. R’s success in the data analysis community stems from two factors described in the epitaphs above: R provides most of the technical power that statisticians require built into the default language, and R has been supported by a community of statisticians who are also open source devotees.
There are many technical advantages afforded by a language
designed specifically for statistical computing. As the description from
the R Project notes, the language provides an open-source bridge to S,
which contains many highly-specialized statistical operations as base
functions. For example, to perform a basic linear regression in R, one
must simply pass the data to the lm
function, which then returns an object containing detailed information
about the regression (coefficients, standard errors, residual values,
etc.). This data can then be visualized by passing the results to the
plot function, which is designed to
visualize the results of this analysis.
In other languages with large scientific computing communities,
such as Python, duplicating the functionality of lm requires the use of several third-party
libraries to represent the data (NumPy), perform the analysis (SciPy)
and visualize the results (matplotlib). As we will see in the
following chapters, such sophisticated analyses can be performed with a
single line of code in R.
In addition, as in other scientific computing environments, the fundamental data type in R is a vector. Vectors can be aggregated and organized in various ways, but at the core, all data are represented this way. This relatively rigid perspective on data structures can be limiting, but is also logical given the application of the language. The most frequently used data structure in R is the data frame, which can be thought of as a matrix with attributes, an internally defined “spreadsheet” structure, or relational database-like structure in the core of the language. Fundamentally, a data frame is simply a column-wise aggregation of vectors that R affords specific functionality to, which makes it ideal for working with any manner of data.
Warning
For all of its power, R also has its disadvantages. R does not scale well with large data, and while there have been many efforts to address this problem, it remains a serious issue. For the purposes of the case studies we will review, however, this will not be an issue. The data sets we will use are relatively small, and all of the systems we will build are prototypes or proof-of-concept models. This distinction is important, because if your intention is to build enterprise level machine learning systems at the Google or Facebook scale, then R is not the right solution. In fact, companies like Google and Facebook often use R as their “data sandbox,” to play with data and experiment with new machine learning methods. If one of those experiments bears fruit, then the engineers will attempt to replicate the functionality designed in R in a more appropriate language, such as C.
This ethos of experimentation has also engendered a great sense of community around the language. The social advantages of R hinge on this large and growing community of experts using and contributing to the language. As Bo Cowgill alludes to, R was borne out of statisticians’ desire to have a computing environment that met their specific needs. Many R users, therefore, are experts in their various fields. This includes an extremely diverse set of disciplines, including mathematics, statistics, biology, chemistry, physics, psychology, economics, and political science, to name a few. This community of experts has built a massive collection of packages on top of the extensive base functions in R. At the time of writing, CRAN contained over 2,800 packages. In the case studies that follow, we will use many of the most popular packages, but this will only scratch the surface of what is possible with R.
Finally, while the latter portion of Cowgill’s statement may seem
a bit menacing, it further highlights the strength of the R community.
As we will see, the R language has a particularly odd syntax that is
rife with coding “gotchas” that can drive even experienced developers
away. But all grammatical grievances with a language can eventually be
overcome, especially for persistent hackers. What is more difficult for
non-statisticians is the liberal assumption of familiarity with
statistical and mathematical methods built into R functions. Using the
lm function as an example, if you had
never performed a linear regression, you would not know to look for
coefficients, standard errors, or residual values in the results. Nor
would you know how to interpret those results.
But, because the language is open source, you are always able to look at the code of a function to see exactly what it is doing. Part of what we will attempt to accomplish with this book is to explore many of these functions in the context of machine learning, but that will ultimately only address a tiny subset of what you can do in R. Fortunately, the R community is full of people willing to help you understand not only the language, but also the methods implemented in it. Table 1-1 lists some of the best places to start.
| Resource | Location | Description |
| RSeek | http://rseek.org/ | When the core development team decided to create an open-source version of S and call it R, they had not considered how hard it would be to search for documents related to a single-letter language on the Web. This specialized search tool attempts to alleviate this by providing a focused portal to R documentation and information. |
| Official R mailing lists | http://www.r-project.org/mail.html | There are several listservs dedicated to the R language, including announcements, packages, development—and of course—help. Many of the language’s core developers frequent these lists, and responses are often quick and terse. |
| StackOverflow | http://stackoverflow.com/questions/tagged/r | Hackers will know StackOverflow.com as one of the premier web resources for coding tips in any language, and the R tag is no exception. Thanks to the efforts of several prominent R community members, there is an active and vibrant collection of experts adding and answering R questions on StackOverflow. |
| #rstats Twtter hash-tag | http://search.twitter.com/search?q=%23rstats | There is also a very active community of R users on Twitter, and they have adopted the #rstats hashtag as their signifier. The thread is a great place to find links to useful resources, find experts in the language, and post questions—as long as they can fit into 140 characters! |
| R-Bloggers | http://www.r-bloggers.com/ | There are hundreds of people blogging about how they use R in their research, work, or just for fun. R-bloggers.com aggregates these blogs and provides a single source for all things related to R in the blogosphere, and is a great place to learn by example. |
| Video Rchive | http://www.vcasmo.com/user/drewconway | As the R community grows, so too do the number of regional meetups and gatherings related to the language. The Rchive attempts to document the presentations and tutorials given at these meetings by posting videos and slides, and now contains presentations from community members all over the world. |
The remainder of this chapter focuses on getting you set up with R and using it. This includes downloading and installing R, as well as installing R packages. We conclude with a miniature case study that will serve as an introduction to some of the R idioms we’ll use in later chapters. This includes issues of loading, cleaning, organizing, and analyzing data.
Downloading and Installing R
Like many open source projects, R is distributed by a series of regional mirrors. If you do not have R already installed on your machine, the first step is to download it. Go to http://cran.r-project.org/mirrors.html and select the CRAN mirror closest to you. Once you have selected a mirror, you will need to download the appropriate distribution of R for whichever operating system you are running.
R relies on several legacy libraries compiled from C and Fortran. As such, depending on your operating system and your familiarity with installing software from source code, you may choose whether to install R from a compiled binary distribution or the source. Below, we present instruction for installing R on Windows, Mac OS X, and Linux distributions, with notes on installing from either source or binaries when available.
Finally, R is available in both 32- and 64-bit versions and, depending on your hardware and operating system combination, you should install the appropriate version.
Windows
For Windows operating systems there are two subdirectories
available to install R: base and
contrib. The latter is a
directory of compiled Windows binary versions of the all of the
contributed R packages in CRAN, while the former is the basic
installation. Select the base
installation, and download the latest compiled binary. Installing
contributed packages is easy to do from R itself and is not
language-specific; therefore, it is not necessary to to install
anything from the contrib
directory. Follow the on-screen instructions for the
installation.
Once the installation has successfully completed, you will have an R application in your Start menu, which will open the RGui and R Console, as pictured in Figure 1-1.
For most standard Windows installations, this process should proceed without any issues. If you have a customized installation, or encounter errors during the installation, consult the R for Windows FAQ at your mirror of choice.
Mac OS X
Fortunately for Mac OS X users, R comes pre-installed with the
operating system. You can check this by opening the Terminal.app and simply typing R at the command-line. You are now ready
to begin! For some users, however, it will be useful to have a GUI
application to interact with the R console. For this you will need
to install separate software. With Mac OS X, you have the option of
installing from either a compiled binary or the source. To install
from a binary—recommended for users with no experience using a Linux command
line—simply download the latest version at your mirror of choice at
http://cran.r-project.org/mirrors.html and
following the on-screen instructions. Once the installation is
complete, you will have both R.app (32-bit) and R64.app (64-bit) available in your
Applications folder. Depending on your version of Mac OS X and your
machine’s hardware, you may choose which version you wish to work
with.
As with the Windows installation, if you are installing from binary this process should proceed without any problems. When you open your new R application you will see a console similar to the one pictured in Figure 1-2.
Note
If you have a custom installation of Mac OS X or wish to customize the installation of R for your particular configuration, we recommend that you install from the source code. To install R from source on Mac OS X requires both C and Fortran compilers, which are not included in the standard installation of the operating system. You can install these compilers using the Mac OS X Developers Tools DVD included with your original Mac OS X installation package, or you can install the necessary compilers from the tools directory at the mirror of your choice.
Once you have all of the necessary compilers to install from
source, the process is the typical configure, make, and install
procedure used to install most software at the command line. Using
the Terminal.app, navigate to the
folder with the source code and execute the following
commands:
$ ./configure $ make $ make install
Depending on your permission settings, you may have to invoke
the sudo command as a prefix to
the configuration step and provide your system password. If you
encounter any errors during the installation, using either the
compiled binary distribution or the source code, consult the
R for Mac OS X FAQ at the mirror
of your choice.
Linux
As with Mac OS X, R comes preinstalled on many Linux
distributions. Simply type R at
the command line and the R console will be loaded. You can now begin
programming! The CRAN mirror also includes installations specific to
several Linux distributions, with instructions for installing R on
Debian, RedHat, SUSE, and Ubuntu. If you use one of these
installations, we recommend that you consult the instructions for
your operating system, because there is considerable variance in the
best practices between Linux distributions.
IDEs and Text Editors
R is a scripting language and therefore the majority of the work done in this book’s case studies will be done within a IDE or text editor, rather than directly inputted into the R console. As we will show in the next section, some tasks are well suited for the console, such as package installation, but primarily you will want to be working within the IDE or text editor of your choice.
For those running the GUI in either Windows or Mac OS X, there
is a basic text editor available from that application. By navigating
to File→New
Document from the menu bar, or clicking on the blank
document icon in the header of the window (highlighted in Figure 1-3), you will open a blank
document in the text editor. As a hacker, you likely already have an
IDE or text editor of choice, and we recommend that you use whichever
environment you are most comfortable in for the case studies. There
are simply too many options to enumerate here, and we have no
intention of inserting ourselves in the infamous emacs versus vim debate.
Loading and Installing R Packages
There are many well-designed, maintained, and supported R
packages related to machine learning. Loading packages in R is very
straightforward. There are two functions to perform this: library and require. There are some subtle differences
between the two, but for the purposes of this book, the primary
difference is that require will
return a Boolean (TRUE or FALSE) value, indicating whether the package
is installed on the machine after attempting to load it. As an
example, below we use library to
load spatstat but require for lda. By using the print function, we can see that we have
lda installed because a Boolean
value of TRUE was returned after
the package was loaded:
library(spatstat) print(require(lda)) [1] TRUE
If we did not have lda
installed (i.e., FALSE was returned
by require), then we would need to
install that package before proceeding.
Note
If you are working with a fresh installation of R, then you will have to install a number of packages to complete all of the case studies in this book.
There are two ways to install packages in R; either with the GUI
interface or with the install.packages function from the console.
Given the intended audience for this book, we will be interacting with
R exclusively from the console during the case studies, but it is
worth pointing out how to use the GUI interface to install packages.
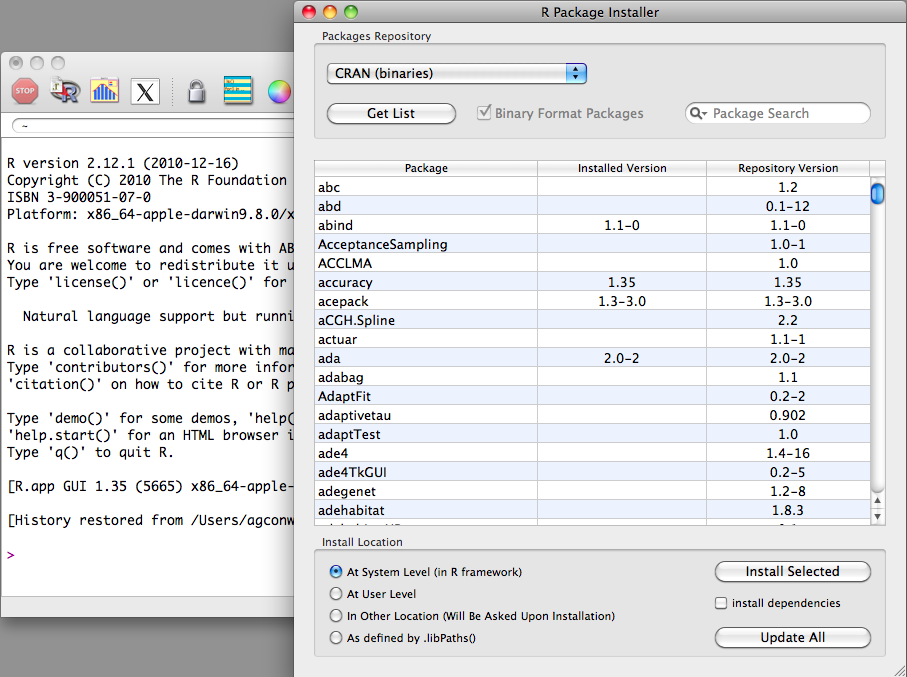
From the menu bar in the application, navigate to Packages &Data→Package Installer, and a window
will appear as displayed in Figure 1-4. From the
Package Repository drop-down,
select either CRAN (binaries) or
CRAN (sources) and click the
Get List button to load all of the
packages available for installation. The most recent version of
packages will be available in the CRAN
(sources) repository, and if you have the necessary
compilers installed on your machine, we recommend using the sources
repository. You can now select the package you wish to install and
click Install Selected to install
the packages.
The install.packages function
is the preferred way to install packages because it provides greater
flexibility in how and where packages get installed. One of the
primary advantages of using install.packages is that it allows you to
install from local source code as well as from CRAN. Though uncommon,
occasionally you may may want to install a package that is not yet
available on CRAN—for example, if you’re updating to an experimental
version of a package. In these cases you will need to install from
source:
install.packages("tm", dependencies=TRUE)
setwd("~/Downloads/")
install.packages("RCurl_1.5-0.tar.gz", repos=NULL, type="source")In the first example above, we use the default settings to
install the tm package from CRAN.
The tm provides function used to do
text mining, and we will use it in Chapter 3
to perform classification on email text. One useful parameter in the
install.packages function is
suggests, which by default is set
to FALSE but if activated will
instruct the function to download and install any secondary packages
used by the primary installation. As a best practice, we recommend
always setting this to TRUE,
especially if you are working with a clean installation of R.
Alternatively, we can also install directly from compressed
source files. In the example above, we are installing the RCurl package from the source code available
on the author’s website. Using the setwd function to make sure the R working
directory is set to the directory where the source file has been
saved, we can simply execute the above command to install directly
from the source code. Note the two parameters that have been altered
in this case. First, we must tell the function not to use one of the
CRAN repositories by setting repos=NULL, and specify the type of
installation using type="source".
As mentioned, we will use several packages through the course of this text. Table 1-2 lists all of the packages used in the case studies and includes a brief description of their purpose, along with a link to additional information about each.
We are now ready to begin exploring machine learning with R! Before we proceed to the case studies, however, we will review some R functions and operations that we will use frequently.
| Name | Location | Author | Description & Use |
ggplot2 | http://had.co.nz/ggplot2/ | Hadley Wickham | An implementation of the grammar of graphics in R. The premier package for creating high-quality graphics. |
plyr | http://had.co.nz/plyr/ | Hadley Wickham | A set of tools used to manipulate, aggregate and manage data in R. |
tm | http://www.spatstat.org/spatstat/ | Ingo Feinerer | A collection of functions for performing text mining in R. Used to work with unstructured text data. |
R Basics for Machine Learning
UFO Sightings in the United States, from 1990-2010
As we stated at the outset, we believe that the best way to learn a new technical skill is to start with a problem you wish to solve or a question you wish to answer. Being excited about the higher level vision of your work makes makes learning from case studies work. In this review of basic concepts in the R language, we will not be addressing a machine learning problem, but we will encounter several issues related to working with data and managing it in R. As we will see in the case studies, quite often we will spend the bulk of our time getting the data formatted and organized in a way that suits the analysis. Very little time, in terms of coding, is usually spent running the analysis.
For this case we will address a question with pure entertainment value. Recently, the data service Infochimps.com released a data set with over 60,000 documented reports of unidentified flying objects (UFO) sightings. The data spans hundreds of years and has reports from all over the world. Though it is international, the majority of sightings in the data come from the United States. With the time and spatial dimensions of the data, one question we might ask is: are there seasonal trends in UFO sightings; and what, if any, variation is there among UFO sightings across the different states in the U.S.?
This is a great data set to start exploring, because it is rich,
well-structured, and fun to work with. It is also useful for this
exercise because it is a large text file, which is typically the type
of data we will deal with in this book. In such text files there are
often messy parts, so we will use base functions in R and some
external libraries to clean and organize the raw data. This section
will take you step-by-step through an entire simple analysis that
tries to answer the questions we posed earlier. You will find the code
for this section in the code folder
for this chapter as the ufo_sightings.R file. We begin by loading
the data and required libraries for the analysis.
Loading libraries and the data
First, we will load the ggplot2 package, which we will use in the
final steps of our visual analysis:
library(ggplot2)
While loading ggplot2, you
will notice that this package also loads two other required
packages: plyr and reshape. Both of these packages are used
for manipulating and organizing data in R, and we will use plyr in this example to aggregate and
organize the data.
The next step is to load the data into R from the text file
ufo_awesome.tsv, which is located
in data/ufo/ directory for this
chapter. Note that the file is tab-delimited (hence the .tsv file extension), which means we will
need to use the read.delim
function to load the data. Because R exploits defaults very heavily,
we have to be particularly conscientious of the default parameter
settings for the functions we use in our scripts. To see how we can
learn about parameters in R, suppose that we had never used the
read.delim function before and
needed to read the help files. Alternatively, assume that we do not
know that read.delim exists and
need to find a function to read delimited data into a data frame. R
offers several useful functions for searching for help:
?read.delim # Access a function's help file
??base::delim # Search for 'delim' in all help files for functions
# in 'base'
help.search("delimited") # Search for 'delimited' in all help files
RSiteSearch("parsing text") # Search for the term 'parsing text' on the R site.In the first example, we append a question mark to the
beginning of the function. This will open the help file for the
given function and it’s an extremely useful R shortcut. We can also
search for specific terms inside of packages by using a combination
of ?? and ::. The double question marks indicate a
search for a specific term. In the example above, we are searching
for occurrences of the term “delim” in all base functions, using the double colon. R
also allows you to perform less structured help searches with
help.search and RSiteSearch. The help.search function will search all help
files in your installed packages for some term, which in the above
example is “delimited.” Alternatively, you can search the R website,
which includes help files and the mailing lists archive, using the
RSiteSearch function. Please
note, this chapter is by no means meant to be an exhaustive review
of R or the functions used in this section. As such, we highly recommend using these search
functions to explore R’s base functions on your own.
For the UFO data there are several parameters in read.delim that we will need to set by
hand in order to properly read in the data. First, we need to tell
the function how the data are delimited. We know this is a
tab-delimited file, so we set sep
to the tab character. Next, when read.delim is reading in data it attempts
to convert each column of data into an R data type using several
heuristics. In our case, all of the columns are strings, but the
default setting for all read.*
functions is to convert strings to factor types. This class is meant for
categorical variables, but we do not want this. As such, we have to
set stringsAsFactors=FALSE to
prevent this. In fact, it is always a good practice to switch off
this default, especially when working with unfamiliar data.
Note
The term “categorical variable” refers to a type of data
that denotes an observation’s membership in a category. In
statistics categorical variables are very important because we may
be interested in what makes certain observations belong to a
certain type. In R we represent categorical variables as factor types, which essentially assigns
numeric references to string labels. In this case, we convert
certain strings—such as state abbreviations—into categorical
variables using as.factor,
which assigns a unique numeric ID to each state abbreviation in
the data set. We will repeat this process many times.
Also, this data does not include a column header as its first
row, so we will need to switch off that default as well to force R
to not use the first row in the data as a header. Finally, there are
many empty elements in the data, and we want to set those to the
special R value NA. To do this we
explicitly define the empty string as the na.string:
ufo<-read.delim("data/ufo/ufo_awesome.tsv", sep="\t", stringsAsFactors=FALSE,
header=FALSE, na.strings="")We now have a data frame containing all of the UFO data!
Whenever working with data frames, especially if they are from
external data sources, it is always a good idea to inspect the data
by hand. Two great functions for doing this are head and tail. These functions will print the first
and last six entries in a data frame:
head(ufo)
V1 V2 V3 V4 V5 V6
1 19951009 19951009 Iowa City, IA <NA> <NA> Man repts. witnessing "flash...
2 19951010 19951011 Milwaukee, WI <NA> 2 min. Man on Hwy 43 SW of Milwauk...
3 19950101 19950103 Shelton, WA <NA> <NA> Telephoned Report:CA woman v...
4 19950510 19950510 Columbia, MO <NA> 2 min. Man repts. son's bizarre sig...
5 19950611 19950614 Seattle, WA <NA> <NA> Anonymous caller repts. sigh...
6 19951025 19951024 Brunswick County, ND <NA> 30 min. Sheriff's office calls to re...The first obvious issue with the data frame is that the column
names are generic. Using the documentation for this data set as a
reference, we can assign more meaningful labels to the columns.
Having meaningful column names for data frames is an important best
practice. It makes your code and output easier to understand, both
for you and other audiences. We will use the names function, which can either access
the column labels for a data structure or assign them. From the data
documentation, we construct a character vector the corresponds to
the appropriate column names and pass it to the names functions with the data frame as its
only argument:
names(ufo)<-c("DateOccurred","DateReported","Location","ShortDescription",
"Duration","LongDescription")From the head output and
the documentation used to create column headings, we know that the
first two columns of data are dates. As in other languages, R treats
dates as a special type, and we will want to convert the date
strings to actual date types. To do this we will use the as.Date function, which will take the date
string and attempt to convert it to as Date object. With this data the strings
have an uncommon date format of the form YYYYMMDD. As such, we will also have to
specify a format string in as.Date so the function knows how to
convert the strings. We begin by converting the DateOccurred column:
ufo$DateOccurred<-as.Date(ufo$DateOccurred, format="%Y%m%d") Error in strptime(x, format, tz = "GMT") : input string is too long
We’ve just come upon our first error! Though a bit cryptic,
the error message contains the substring “input string too long”,
which indicates that some of the entries in the DateOccurred column are too long to match
the format string we provided. Why might this be the case? We are
dealing with a large text file, so perhaps some of the data was
malformed in the original set. Assuming this is the case, those data
points will not be parsed correctly when being loaded by read.delim and that would cause this sort
of error. Because we are dealing with real world data, we’ll need to
do some cleaning by hand.
Converting date strings, and dealing with malformed data
To address this problem we first need to locate the rows with
defective date strings, then decide what to do with them. We are
fortunate in this case because we know from the error that the
errant entries are “too long.” Properly parsed strings will always
be eight characters long, i.e., “YYYYMMDDD”. To find the problem
rows, therefore, we simply need to find those that have strings with
more than eight characters. As a best practice, we first inspect the
data to see what the malformed data looks like, in order to get a
better understanding of what has gone wrong. In this case, we will
use the head function, as before,
to examine the data returned by our logical statement.
Later, to remove these errant rows we will use the ifelse function to construct a vector of
TRUE and FALSE values to identify the entries that
are eight characters long (TRUE)
and those that are not (FALSE).
This function is a vectorized version of the typical if-else logical
switch for some Boolean test. We will see many examples of
vectorized operations in R. They are the preferred mechanism for
iterating over data because they are often—but not always—more
efficient:[3]
head(ufo[which(nchar(ufo$DateOccurred)!=8 | nchar(ufo$DateReported)!=8),1])
[1] "ler@gnv.ifas.ufl.edu"
[2] "0000"
[3] "Callers report sighting a number of soft white balls of lights headingin
an easterly directing then changing direction to the west beforespeeding off to
the north west."
[4] "0000"
[5] "0000"
[6] "0000"
good.rows<-ifelse(nchar(ufo$DateOccurred)>!=8 | nchar(ufo$DateReported)!=8,
FALSE,TRUE)
length(which(!good.rows))
[1] 371
ufo<-ufo[good.rows,]We use several useful R functions to perform this search. We
need to know the length of the string in each entry of DateOccurred and DateReported, so we use the nchar function to compute this. If that
length is not equal to eight, then we return FALSE. Once we have the vectors of
Booleans, we want to see how many entries in the data frame have
been malformed. To do this, we use the which command to return a vector of vector
indices that are FALSE. Next, we
compute the length of that vector
to find the number of bad entries. With only 371 rows not
conforming, the best option is to simply remove these entries and
ignore them. At first, we might worry that losing 371 rows of data
is a bad idea, but there are over sixty-thousand total rows, so we
will simply ignore those rows and continue with the conversion to
Date types:
ufo$DateOccurred<-as.Date(ufo$DateOccurred, format="%Y%m%d")
ufo$DateReported<-as.Date(ufo$DateReported, format="%Y%m%d")Next we will need to clean and organize the location data.
Recall from the previous head
call that the entries for UFO sightings in the United States take
the form “City, State”. We can use R’s regular expression
integration to split these strings into separate columns and
identify those entries that do not conform. The latter portion
(identifying those that do not conform) is particularly important
because we are only interested in sighting variation in the United
States and will use this information to isolate those
entries.
Organizing location data
To manipulate the data in this way, we will first construct a
function that takes a string as input and performs the data
cleaning. Then we will run this function over the location data
using one of the vectorized apply
functions:
get.location<-function(l) {
split.location<-tryCatch(strsplit(l,",")[[1]], error= function(e) return(c(NA, NA)))
clean.location<-gsub("^ ","",split.location)
if (length(clean.location)>2) {
return(c(NA,NA))
}
else {
return(clean.location)
}
}There are several subtle things happening in this function.
First, notice that we are wrapping the strsplit command in R’s error handling
function, tryCatch. Again, not
all of the entries are of the proper “City, State” form, and in fact
some do not even contain a comma. The strsplit function will throw an error if
the split character is not matched, therefore, we have to catch this
error. In our case, when there is no comma to split we will return a
vector of NA to indicate that
this entry is not valid. Next, the original data included leading
whitespace, so we will use the gsub function (part of R’s suite of
functions for working with regular expressions) to remove the
leading whitespace from each character. Finally, we add an
additional check to ensure that only the location vectors of length
two are returned. Many non-U.S. entries have multiple commas,
creating larger vectors from the strsplit function. In this case, we will
again return an NA vector.
With the function defined, we will use the lapply function, short for “list-apply,”
to iterate this function over all strings in the Location column. As mentioned, the
apply family of functions in R
are extremely useful. They are constructed of the form apply(vector, function), and return
results of the vectorized application of the function to the vector
in a specific form. In our case, we are using lapply, which always returns a list:
city.state<-lapply(ufo$Location, get.location) head(city.state) [[1]] [1] "Iowa City" "IA" [[2]] [1] "Milwaukee" "WI" [[3]] [1] "Shelton" "WA" [[4]] [1] "Columbia" "MO" [[5]] [1] "Seattle" "WA" [[6]] [1] "Brunswick County" "ND"
From above, a list in R is
a key-value style data structures, wherein the keys are indexed by
the double-bracket and values by the single bracket. In our case the
keys are simply integers, but lists can also have strings as
keys.[4] Though convenient, having the data stored in a
list is not desirable, as we
would like to add the city and state information to the data frame
as separate columns. To do this we will need to convert this long
list into a two-column matrix, with the city data as the leading
column:
location.matrix<-do.call(rbind, city.state)
ufo<-transform(ufo, USCity=location.matrix[,1], USState=tolower(location.matrix[,2]),
stringsAsFactors=FALSE)To construct a matrix from the list, we use the do.call function. Similar to the apply functions, do.call executes a function call over a
list. We will often use the combination of lapply and do.call to manipulate data. Above we pass
the rbind function, which will
“row-bind” all of the vectors in the city.state list to create a matrix. To get
this into the data frame we use the transform function. We create two new
columns; USCity and USState, from the first and second columns
of location.matrix, respectively.
Finally, the state abbreviations are inconsistent, with some
uppercase and others lowercase, so we use the tolower function to make them all
lowercase.
Dealing with data outside our scope
The final issue related to data cleaning we must consider
concerns entries that meet the “City, State” form, but are not from
the U.S. Specifically, the data include several UFO sightings from
Canada, which also take this form. Fortunately, none of the Canadian
province abbreviations match U.S. state abbreviations. We can use
this information to identify non-U.S. entries by constructing a
vector of U.S. state abbreviations and only keeping those entries in
the USState column that match an
entry in this vector:
us.states<-c("ak","al","ar","az","ca","co","ct","de","fl","ga","hi","ia","id","il",
"in","ks","ky","la","ma","md","me","mi","mn","mo","ms","mt","nc","nd","ne","nh",
"nj","nm","nv","ny","oh","ok","or","pa","ri","sc","sd","tn","tx","ut","va","vt",
"wa","wi","wv","wy")
ufo$USState<-us.states[match(ufo$USState,us.states)]
ufo$USCity[is.na(ufo$USState)]<-NATo find the entries in the USState column that do not match a U.S.
state abbreviation, we use the match function. This function takes two
arguments: the first are the values to be matched, and the second
those to be matched against. What is returned is a vector of the
same length as the first argument, in which the values are the index
of entries in that vector that match some value in the second
vector. If no match is found, the function returns NA by default. In our case, we are only
interested in which entries are NA, as these are those entries that do not
match a state. We then use the is.na function to find which entries are
not U.S. states and reset them to NA in the USState column. Finally, we also set those
indices in the USCity column to
NA for consistency.
Our original data frame now has been manipulated to the point
that we can extract from it only the data we are interested in.
Specifically, we want a subset that includes only U.S. incidents of
UFO sightings. By replacing entries that did not meet this criteria
in the previous steps, we can use the subset command to create a new data frame
of only U.S. incidents:
ufo.us<-subset(ufo, !is.na(USState)) head(ufo.us) DateOccurred DateReported Location ShortDescription Duration 1 1995-10-09 1995-10-09 Iowa City, IA <NA> <NA> 2 1995-10-10 1995-10-11 Milwaukee, WI <NA> 2 min. 3 1995-01-01 1995-01-03 Shelton, WA <NA> <NA> 4 1995-05-10 1995-05-10 Columbia, MO <NA> 2 min. 5 1995-06-11 1995-06-14 Seattle, WA <NA> <NA> 6 1995-10-25 1995-10-24 Brunswick County, ND <NA> 30 min. LongDescription USCity USState 1 Man repts. witnessing "flash... Iowa City ia 2 Man on Hwy 43 SW of Milwauk... Milwaukee wi 3 Telephoned Report:CA woman v... Shelton wa 4 Man repts. son's bizarre sig... Columbia mo 5 Anonymous caller repts. sigh... Seattle wa 6 Sheriff's office calls to re... Brunswick County nd
Aggregating and organizing the data
We now have our data in a format in which we can begin
analyzing it! In the previous section we spent a lot of time getting
the data properly formatted and identifying the relevant entries for
our analysis. In this section we will explore the data to further
narrow our focus. These data have two primary dimensions: space
(where the sighting happened) and time (when a sighting occurred).
We focused on the former in the previous section, but here we will
focus on the latter. First, we use the summary function on the DateOccurred column to get a sense of this
chronological range of the data:
summary(ufo.us$DateOccurred) Min. 1st Qu. Median Mean 3rd Qu. Max. "1400-06-30" "1999-09-06" "2004-01-10" "2001-02-13" "2007-07-26" "2010-08-30"
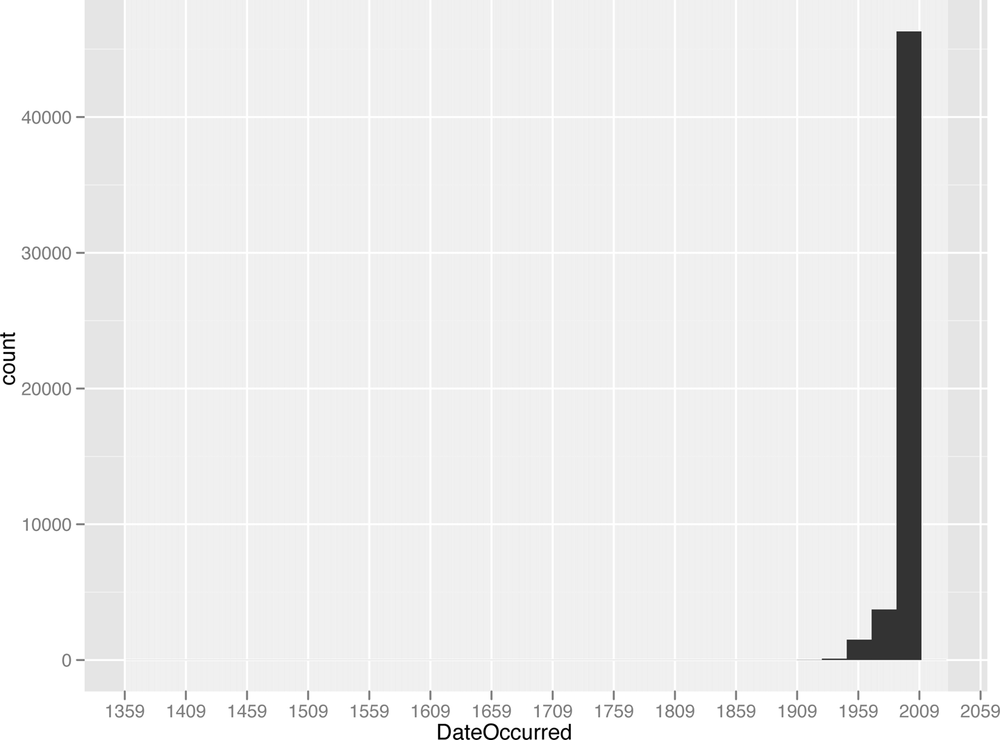
Surprisingly, this data goes back quite a long time:the oldest UFO sighting comes from 1400! Given this outlier, the next question is: how are these data distributed over time? And is it worth analyzing the entire time series? A quick way to look at this visually is to construct a histogram. We will discuss histograms in more detail in the next chapter, but for now you should know that histograms allow you to bin your data by a given dimension and observe the frequency with which your data falls into those bins. The dimension of interest here is time, so we construct a histogram that bins the data over time:
quick.hist<-ggplot(ufo.us, aes(x=DateOccurred))+geom_histogram()+
scale_x_date(major="50 years")
ggsave(plot=quick.hist, filename="../images/quick_hist.png", height=6, width=8)
stat_bin: binwidth defaulted to range/30. Use 'binwidth = x' to adjust this.There are several things to note here. This is our first use
of the ggplot2 package, which we
will use throughout the book for all of our data visualizations. In
this case, we are constructing a very simple histogram, which we
only requires a single line of code. First, we create a ggplot object and pass it the UFO data
frame as its initial argument. Next, we set the x-axis aesthetic to
the DateOccurred column, as this
is the frequency we are interested in examining. With ggplot2 we must always work with data
frames, and the first argument to create a ggplot object must always be a data frame.
ggplot2 is an R implementation of
Leland Wilkinson’s Grammar of Graphics
Wil05. This means the package adheres to this
particular philosophy for data visualization, and all visualizations
will be built up as a series of layers. For this histogram, the
initial layer is the x-axis data, namely the UFO sighting dates.
Next, we add an histogram layer with the geom_histogram function. In this case, we
will use the default settings for this function, but, as we will see
later, this default is often not a good choice. Finally, because
this data spans such a long time period, we will rescale the x-axis
labels to occur every 50 years with the scale_x_date function.
Once the ggplot object has
been constructed, we use the ggsave function to output the
visualization to a file. We could have also used > print(quick.hist) to print the
visualization to the screen. Note the warning message that is
printed when you draw the visualization. There are many ways to bin
data in a histogram, and we will discuss this in detail the next
chapter, but this warning is provided to let you know exactly how
ggplot2 does the binning by
default.
We are now ready to explore the data with this visualization, which is illustrated in Figure 1-5.
The results of this analysis are stark. The vast majority of
the data occur between 1960 and 2010, with the majority of UFO
sightings occurring within the last two decades. For our purposes,
therefore, we will focus on only those sightings that occurred
between 1990 and 2010. This will allow us to exclude the outliers
and compare relatively comparable units during the analysis. As
before, we will used the subset
function to create a new data frame that meets this criteria:
ufo.us<-subset(ufo.us, DateOccurred>=as.Date("1990-01-01"))
nrow(ufo.us)
[1] 46347While this removes many more entries than we eliminated while
cleaning the data, it still leaves us with over forty-six thousand
observations to analyze. Next, we must begin organizing the data
such that it can be used to address our central question: what, if
any, seasonal variation exists for UFO sightings in U.S. states? To
address this, we must first ask: what do we mean by “seasonal?”
There are many ways to aggregate time series data with respect to
seasons; by week, month, quarter, year, etc. But which way of
aggregating our data is most appropriate here? The DateOccurred column provides UFO sighting
information by the day, but there is considerable inconsistency in
terms of the coverage throughout the entire set. We need to
aggregate the data in a way that puts the amount of data for each
state on relatively level planes. In this case, doing so by
year-month is the best option. This aggregation also best addresses
the core of our question, as monthly aggregation will give good
insight into seasonal variations.
We need to count the number of UFO sightings that occurred in
each state by all year-month combinations from 1990-2010. First, we
will need to create a new column in the data that corresponds to the
year and months present in the data. We will use the strftime function to convert the Date objects to a string of the “YYYY-MM”
format. As before, we will set the format parameter accordingly to get the
strings:
ufo.us$YearMonth<-strftime(ufo.us$DateOccurred, format="%Y-%m")
Notice that in this case we did not use the transform function to add a new column to
the data frame. Rather, we simply referenced a column name that did
not exists and R automatically added it. Both methods for adding new
columns to a data frame are useful, and we will switch between them
depending on the particular task. Next, we want to count the number
of times each state and year-month combination occurs in the data.
For the first time we will use the ddply function, which is part of the
extremely useful plyr library for
manipulating data.
The plyr family of
functions work a bit like the map-reduce style data aggregation
tools that have risen in popularity over the past several years.
They attempt to group data in some specific way meaningful to all
observations, and then do some calculation on each of these group
and return the results. For this task we want to group the data by
state abbreviations and the year-month column we just created. Once
the data is grouped as such, we count the number of entries in each
group and return that as a new column. Here we will simply use the
nrow function to reduce the data
by the number of rows in each group:
sightings.counts<-ddply(ufo.us,.(USState,YearMonth), nrow) head(sightings.counts) USState YearMonth V1 1 ak 1990-01 1 2 ak 1990-03 1 3 ak 1990-05 1 4 ak 1993-11 1 5 ak 1994-11 1 6 ak 1995-01 1
We now have the number of UFO sightings for each state by the
year and month. From the head
call above, however, we can see that there may be a problem with
using the data as is, because it contains a lot of missing values.
For example, we see that there was one UFO sighting in January,
March, and May of 1990 in Arkansas; but no entries appear for
February or April. Presumably, there were no UFO sightings in these
months, but the data does not include entries for non-sightings, so
we have to go back and add these as zeroes.
We need a vector of years and months that span the entire data
set. From this we can check to see if they are already in the data,
and if not, add them as zeroes. To do this, we will create a
sequence of dates using the seq.Date function, and then format them to
match the data in our data frame:
date.range<-seq.Date(from=as.Date(min(ufo.us$DateOccurred)),
to=as.Date(max(ufo.us$DateOccurred)), by="month")
date.strings<-strftime(date.range, "%Y-%m")With the new date.strings
vector, we need to create a new data frame that has all year-months
and states. We will use this to performing the matching with the UFO
sighting data. As before, we will use the lapply function to create the columns and
the do.call function to convert
this to a matrix and then data frame:
states.dates<-lapply(us.states,function(s) cbind(s,date.strings)) states.dates<-data.frame(do.call(rbind, states.dates), stringsAsFactors=FALSE) head(states.dates) s date.strings 1 ak 1990-01 2 ak 1990-02 3 ak 1990-03 4 ak 1990-04 5 ak 1990-05 6 ak 1990-06
The states.dates data frame
now contains entries for every year, month and state combination
possible in the data. Note that there are now entries for February
and April 1990 for Arkansas. To add in the missing zeroes to the UFO
sighting data, we need to merge this data with our original data
frame. To do this, we will use the merge function, which takes two ordered
data frames and attempts to merge them by common columns. In our
case, we have two data frames ordered alphabetically by U.S. state
abbreviations and chronologically by year and month. We need to tell
the function which columns to merge these data frames by. We will
set the by.x and by.y parameters according to the matching
column names in each data frame. Finally, we set the all parameter to TRUE, which instructs the function to
include entries that do not match and fill them with NA. Those entries in the V1 column will be those state, year, and
month entries for which no UFOs were sighted:
all.sightings<-merge(states.dates,sightings.counts,by.x=c("s","date.strings"),
by.y=c("USState","YearMonth"),all=TRUE)
head(all.sightings)
s date.strings V1
1 ak 1990-01 1
2 ak 1990-02 NA
3 ak 1990-03 1
4 ak 1990-04 NA
5 ak 1990-05 1
6 ak 1990-06 NAThe final steps for data aggregation are simple housekeeping.
First, we will set the column names in the new all.sightings data frame to something
meaningful. This is done in exactly the same way as we did it at the
outset. Next, we will convert the NA entries to zeroes using the is.na function, again. Finally, we will
convert the YearMonth and
State columns to the appropriate
types. Using the date.range
vector we created in the previous step and the rep function to create a new vector that
repeats a given vector, we replace the year and month strings with
the appropriate Date object.
Again, it is better to keep dates as Date objects rather than strings because
we can compare Date objects
mathematically, but we can’t do that easily with strings. Likewise,
the state abbreviations are better represented as a categorical
variables than strings, so we convert these to factor types. We will describe factors, and other R data types, in more
detail in the next chapter:
names(all.sightings)<-c("State","YearMonth","Sightings")
all.sightings$Sightings[is.na(all.sightings$Sightings)]<-0
all.sightings$YearMonth<-as.Date(rep(date.range,length(us.states)))
all.sightings$State<-as.factor(toupper(all.sightings$State))We are now ready to analyze the data visually!
Analyzing the data
For this data we will only address the core question by
analyzing it visually. For the remainder of the book we will combine
both numeric and visual analyses, but as this example is only meant
to introduce core R programming paradigms, we will stop at the
visual component. Unlike the previous histogram visualization,
however, we will take greater care with ggplot2 to build the visual layers
explicitly. This will allow us to create a visualization that
directly addresses the question of seasonal variation among states
overtime and produce a more professional-looking
visualization.
We will construct the visualization all at once below, then explain each layer individually:
state.plot<-ggplot(all.sightings, aes(x=YearMonth,y=Sightings))+geom_line(aes(color="darkblue"))+
facet_wrap(~State,nrow=10,ncol=5)+
theme_bw()+
scale_color_manual(values=c("darkblue"="darkblue"),legend=FALSE)+
scale_x_date(major="5 years", format="%Y")+
xlab("Time")+ylab("Number of Sightings")+
opts(title="Number of UFO sightings by Month-Year and U.S. State (1990-2010)")
ggsave(plot=state.plot, filename="../images/ufo_sightings.pdf",width=14,height=8.5)As always, the first step is to create a ggplot object with a data frame as its
first argument. Here, we are using the all.sightings data frame we created in the
previous step. Again, we need to build an aesthetic layer of data to
plot, and in this case the x-axis is the YearMonth column and the y-axis is the
Sightings data. Next, to show
seasonal variation among states, we will plot a line for each state.
This will allow us to observe any spikes, lulls, or oscillation in
the number of UFO sightings for each state over time. To do this, we
will use the geom_line function
and set the color to “darkblue”
to make the visualization easier to read.
As we have seen throughout this case, the UFO data is fairly
rich and includes many sightings across the United States over a
long period of time. Knowing this, we need to think of a way to
break up this visualization such that we can observe the data for
each state, but also compare it to the others. If we plot all of the
data in a single panel it will be very difficult to discern
variation. To check this, run the first line of code from the above
block, but replace color="darkblue" with color=State and enter > print(state.plot) at the console. A
better approach would be to plot the data for each state
individually, and order them in a grid for easy comparison.
To create a multi-faceted plot, we use the facet_wrap function and specify that the
panels be created by the State
variable, which is already a factor type, i.e., categorical. We also
explicitly define the number of rows and columns in the grid, which
is easier in our case because we know we are creating 50 different
plots.
The ggplot2 package has
many plotting themes. The default theme is the one we used in the
first example and has a grey background with dark grey gridlines.
While it is strictly a matter of taste, we prefer using a white
background for this plot, since that will make it easier to see
slight difference among data points in our visualization. We add the
theme_bw layer, which will
produce a plot with a white background and black gridlines. Once you
become more comfortable with ggplot2, we recommend experimenting with
different defaults to find the one you prefer.[5]
The remaining layers are done as housekeeping to make sure the
visualization has a professional look and feel. Though not formally
required, paying attention to these details is what can separate
amateurish plots from professional-looking data visualizations. The
scale_color_manual function is
used to specify that the string “darkblue” corresponds to the
web-safe color “darkblue.” While this may seem repetitive, it is at
the core of ggplot2’s design,
which requires explicit definition of details, such as color. In
fact, ggplot2 tends to think of
colors as a way of distinguishing among different types or
categories of data; and, as such, prefers to have a factor type used to specify color. In our
case we are defining a color explicitly using a string and therefore
have to define the value of that string with the scale_color_manual function.
As we did before, we use the scale_x_date to specify the major
gridlines in the visualization. Since this data spans twenty years,
we will set these to be at regular five year intervals. Then we set
the tick labels to be the year in a full four digit format. Next, we
set the x-axis label to “Time” and the y-axis label to “Number of
Sightings” by using the xlab and
ylab functions respectively.
Finally, we use the opts function
to give the plot a title.There are many more options available in
the opts function, and in later
chapters we will see some of them, but there are many more beyond
the scope of this book.
With all of the layers built, we are now ready to render the
image with ggsave and analyze the
data. The results are shown in Figure 1-6.
There are many interesting observations that arise in this analysis. We see that California and Washington are large outliers in terms of the number of UFO sightings reported in these states compared to the others. Between these outliers there are also interesting differences. In California, the number of UFO sightings seems to be somewhat random over time, but steadily increasing since 1995; while in Washington, the seasonal variation seems to be very consistent over time, with regular peaks and valleys in UFO sightings starting from about 1995.
We can also notice that many states experience sudden spikes in the number of UFO sightings reported. For example, Arizona, Florida, Illinois, and Montana seem to have experienced spikes around mid-1997, while Michigan, Ohio, and Oregon experienced similar spikes in late 1999. Only Michigan and Ohio are geographically close among these groups. If we do not believe that these are actually the result of extraterrestrial visitors, what are some alternative explanations? Perhaps there was increased vigilance among citizens to look to the sky as the millennium came to a close, causing heavier reporting of false sightings.
If, however, you are sympathetic to the notion that we may be regularly hosting visitors from outer space, there is also evidence to pique your curiosity. In fact, there is surprising regularity of these sightings in many states in the United States, with evidence of regional clustering as well. It is almost as if the sightings really contain a meaningful pattern.
Further Reading on R
This introductory case is by no means meant to be an exhaustive review of the language. Rather, we used this data set to introduce several R paradigms related to loading, cleaning, organizing, and analyzing data. We will revisit many of the functions and processes reviewed above in the following chapters, along with many others. For those readers interested in gaining more practice and familiarity with R before proceeding, there are many excellent resources. These resources can roughly be divided into either reference books and texts or online resources, as shown in Table 1-3.
In the next chapter, we will review exploratory data analysis. Much of the above case study involved exploring data, but we moved through these steps rather quickly. In the next section we will consider the process of data exploration much more deliberately.
| Title | Author | Reference | Description |
| Text References | |||
| Data Manipulation with R | Phil Spector | Spe08 | A deeper review of many of the data manipulation topics covered in the previous section, and introduction to several techniques not covered. |
| R in a Nutshell | Joseph Adler | Adl10 | A detailed exploration of all of R’s base functions. This book takes the R manual and adds several practical examples. |
| Introduction to Scientific Programming and Simulation Using R | Owen Jones, Robert Maillardet, and Andrew Robinson | JMR09 | Unlike other introductory texts to R, this book focuses on the primacy of learning the language first, then creating simulations. |
| Data Analysis Using Regression and Multilevel/Hierarchical Models | Andrew Gelman and Jennifer Hill | GH06 | This text is heavily focused on doing statistical analyses, but all of the examples are in R and is an excellent resources for both learning the language and methods. |
| ggplot2: Elegant Graphics for Data Analysis | Hadley Wickham | Wic09 | The definitive guide to creating data visualizations with ggplot2. |
| Online References | |||
| An Introduction to R | Bill Venables and David Smith | http://lib.stat.cmu.edu/S/Spoetry/Tutor/R_inferno.pdf | An extensive and ever-changing introduction to the language from the R Core team. |
| The R Inferno | Patrick Burns | http://cran.r-project.org/doc/manuals/R-intro.html | An excellent introduction to R for the experienced programmer. The abstract says it best, “If you are using R and you think you’re in hell, this is a map for you.” |
| R for Programmers | Norman Matloff | http://heather.cs.ucdavis.edu/~matloff/R/RProg.pdf | Similar to the “R Inferno,” this introduction is geared for programmers with experience in other languages. |
| The split-apply-combine strategy for data analysis | Hadley Wickham | http://had.co.nz/plyr/plyr-intro-090510.pdf | The author of plyr and provides an excellent
introduction to the map-reduce paradigm in the context of his
tools, with many examples. |
| R Data Analysis Examples | UCLA ATS | http://www.ats.ucla.edu/stat/r/dae/default.htm | A great “Rosetta Stone” style introduction to those with experience in other statistical programming platforms, such as SAS, SPSS and Stata. |
Get Machine Learning for Email now with the O’Reilly learning platform.
O’Reilly members experience books, live events, courses curated by job role, and more from O’Reilly and nearly 200 top publishers.